- Cart 0
- English
Technical Principles and Clinical Detection Applications of Viral RNA Extraction Kits
May 11, 2026
Clicks:71
In molecular diagnostics and virological research, efficient extraction of viral RNA from clinical samples is critical to ensuring the sensitivity and specificity of downstream detection. Viral RNA Isolation Kits are specially designed for liquid samples such as plasma, serum, cell‑free body fluids and cell culture supernatants. Using optimized lysis systems and silica‑membrane‑based nucleic acid binding technology, they enable organic‑solvent‑free nucleic acid purification at room temperature throughout the workflow. This standardized extraction protocol not only shortens experimental time but also addresses the technical challenge of easy RNA degradation through strict RNase‑free environment control.
Why is Viral RNA Extraction Specially Optimized?
Compared with total RNA extraction from cells or tissues, viral RNA extraction faces unique technical challenges: Low abundance and dilution: Viral particle concentrations in plasma or serum are usually extremely low (e.g., HIV‑1 RNA loads can be as low as 50 copies/mL), and sample volume is limited (typically 200 μL), requiring ultra‑high recovery efficiency. RNA fragility: Genomes of RNA viruses are usually single‑stranded, more unstable than double‑stranded DNA. Plasma contains abundant RNases that rapidly degrade RNA without timely treatment after sampling. Therefore, an RNase‑Free environment is not merely a selling point of kits but an essential prerequisite for success. Inhibitor removal: Blood samples contain PCR inhibitors such as hemoglobin, immunoglobulins and lipids. Incomplete removal leads to Ct‑value shifts or even false‑negative results in subsequent RT‑qPCR.
Why Is Room‑Temperature‑Only Operation Feasible?
Traditional RNA extraction relies on ice‑bath lysis and low‑temperature centrifugation (4 °C) to inhibit RNase activity and RNA degradation. However, innovations in chemical formulation enable 15–25 °C room‑temperature‑only operation with Viral RNA Isolation Kits: Instant inactivation mechanism: Lysis Buffer viRL contains high concentrations of chaotropic salts that rapidly disrupt viral envelopes and protein coats while denaturing RNases, eliminating the need for low‑temperature enzyme inhibition. Chemical stability: Silica membranes in spin columns maintain stable nucleic acid binding capacity at room temperature. Combined with isopropanol precipitation, RNA can be efficiently captured without low‑temperature centrifugation. Carrier‑assisted technology: Linear Acrylamide serves as a nucleic acid carrier, co‑precipitating with RNA in the presence of isopropanol to significantly improve recovery of low‑abundance RNA, especially trace viral genomes.
Core Steps of Extraction Workflow
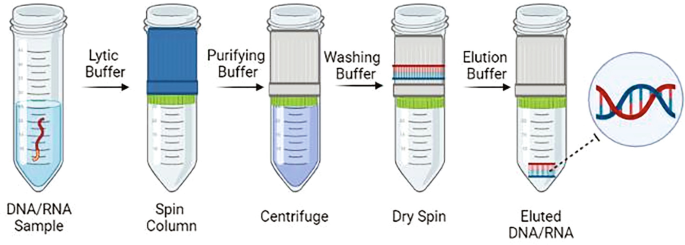
The standard protocol can be completed within 30 min, with elaborately designed key steps: Lysis and carrier binding: Add lysis buffer (containing SDS or chaotropic salts) and Linear Acrylamide to samples, vortex thoroughly to fully lyse viral particles and release RNA. Carrier molecules form complexes with RNA to prevent loss in subsequent steps. Isopropanol‑mediated binding adjustment: Add isopropanol (typically 350 μL) to reduce solution dielectric constant, promoting efficient adsorption of RNA‑carrier complexes onto silica membranes. Centrifuge (8000 rpm, 1 min) to pass liquid through spin columns; RNA is retained on membranes while impurities such as proteins and polysaccharides flow through. Dual‑buffer washing system: Buffer viRW1 (containing chaotropic salts): Removes protein contamination and residual hemolytic products; Buffer viRW2 (containing ethanol): Double washing thoroughly removes salt ions to prevent PCR inhibition. Perform a final dry‑spin centrifugation (12,000 rpm, 2 min) to eliminate residual ethanol that may interfere with downstream reverse transcription. Optimized thermal elution: Elute with 65 °C pre‑warmed RNase‑free water (30–50 μL). Pre‑heating weakens nucleic acid‑membrane binding to improve elution efficiency. If low RNA yield is expected, reload eluate onto the column membrane for repeated elution to maximize recovery.
Which Experiments Depend on High‑Quality Viral RNA?
RNA purity (OD260/280 usually 1.8–2.1) directly determines reliability of downstream applications: Real‑time quantitative PCR (RT‑qPCR): The primary application of viral RNA extraction. High‑purity RNA ensures good linearity of standard curves and accurate low‑background Ct values reflecting viral loads, which is critical for quantitative diagnosis of COVID‑19, HIV, HCV, influenza virus, etc. Next‑generation sequencing (NGS): Viral whole‑genome sequencing or metagenomic studies require intact non‑degraded RNA fragments. High‑quality extracts can be directly used for library construction without rRNA contamination interference. Reverse‑transcription loop‑mediated isothermal amplification (RT‑LAMP): In POCT (point‑of‑care testing) scenarios, rapidly extracted viral RNA can be directly used for isothermal amplification, achieving sample‑to‑result conversion within 30 min. Infectious clone construction: Viral genomic RNA extracted from culture supernatants can be reverse‑transcribed into cDNA for full‑length genomic cloning and reverse genetics studies of viral pathogenic mechanisms. Viral mutation monitoring and drug‑resistance analysis: RT‑PCR and sequencing using extracted RNA monitor viral gene mutations (e.g., SARS‑CoV‑2 variant tracking) and drug‑resistant loci.
How to Avoid Common Operational Pitfalls?
Sample freeze‑thaw risks: Plasma or serum samples should not undergo more than one freeze‑thaw cycle to prevent RNA degradation and yield reduction. For long‑term storage, store directly at −80 °C rather than −20 °C. Precipitation interference: Transfer flocculent precipitates (usually caused by excess proteins or lipids) generated after lysis together to spin columns to avoid discarding viral RNA with precipitates. Increase centrifugation time or speed for excessive precipitates. Critical ethanol washing step: Absolute ethanol must be added to Buffer viRW2 before use (corresponding volume added to 60 mL buffer), otherwise effective desalting and nucleic acid purification cannot be achieved. Elution volume optimization: Elution volume should not be less than 30 μL; insufficient volume fails to fully wet membrane surfaces and causes incomplete elution. Excessively large volume (>50 μL) dilutes RNA concentration and reduces detection sensitivity for low‑copy templates. Cross‑contamination prevention: RNA‑Only spin columns are single‑use and non‑reusable. When processing large batches of samples, set negative (no‑template) and positive controls per batch to monitor contamination and extraction efficiency.
As infectious disease diagnosis and virological research increasingly rely on molecular techniques, standardized viral RNA extraction serves not only as the starting point of experimental workflows but also as the cornerstone of data quality. By understanding the chemical principles and operational details of extraction kits, researchers can rapidly and stably obtain high‑quality viral RNA under biosafety‑compliant conditions, supporting precise diagnosis and fundamental scientific research.
Recommended Absin Viral RNA Isolation Kit
| Cat. No. | Product Name | Size |
|---|---|---|
| abs60355 | Viral RNA Isolation Kit | 50 T/200 T |
Contact Absin
Absin provides antibodies, proteins, ELISA kits, cell culture, detection kits, and other research reagents. If you have any product needs, please contact us.
| Absin Bioscience Inc. worldwide@absin.cn |
 Follow us on Facebook: Absin Bio Follow us on Facebook: Absin Bio |
