- Cart 0
- English
Why is your research on transcription factors always falling short of the final breakthrough?
May 08, 2026
Clicks:69
In the study of gene expression regulation, a question has plagued countless researchers: we clearly know that a transcription factor is involved in a specific signaling pathway and can even observe its regulation of downstream gene expression, but we can never directly prove whether it actually "touches" the regulatory regions of those genes. This dilemma of "seeing the effect but failing to capture the evidence" often stems from the lack of a key technical tool —— Chromatin Immunoprecipitation.
What Exactly is a Chromatin Immunoprecipitation Kit?
Chromatin Immunoprecipitation (ChIP) kit is a complete experimental system dedicated to studying protein-DNA interactions. It uses chemical cross-linking to "lock" intracellular chromatin-binding proteins (mainly transcription factors, histone-modifying enzymes, etc.) with their interacting DNA fragments, then captures the target protein using specific antibodies, and finally co-precipitates the bound DNA sequences.
In terms of components, a complete ChIP kit usually contains the following core modules:
Cross-linking & Lysis System Contains cross-linking agents such as formaldehyde and cell lysis buffer, used to fix protein-DNA complexes and disrupt cell membranes
Chromatin Fragmentation Tools Fragment chromatin into 200-1000 bp segments via sonication or enzymatic digestion
Immunoprecipitation Module Protein A/G magnetic beads are the core of this module, efficiently capturing antibody-antigen complexes
Washing System Various washing buffers (low-salt, high-salt, LiCl, etc.) to remove non-specific binding
DNA Purification Components Reverse cross-linking, proteinase K digestion, and spin column purification to obtain high-purity DNA samples
Detection Supporting Reagents Including positive control antibody (e.g., RNA polymerase II antibody), negative control IgG, and control primers
Such kits usually adopt the magnetic bead method instead of traditional agarose beads, because magnetic beads have more uniform particle size, faster magnetic response, and lower non-specific adsorption background.
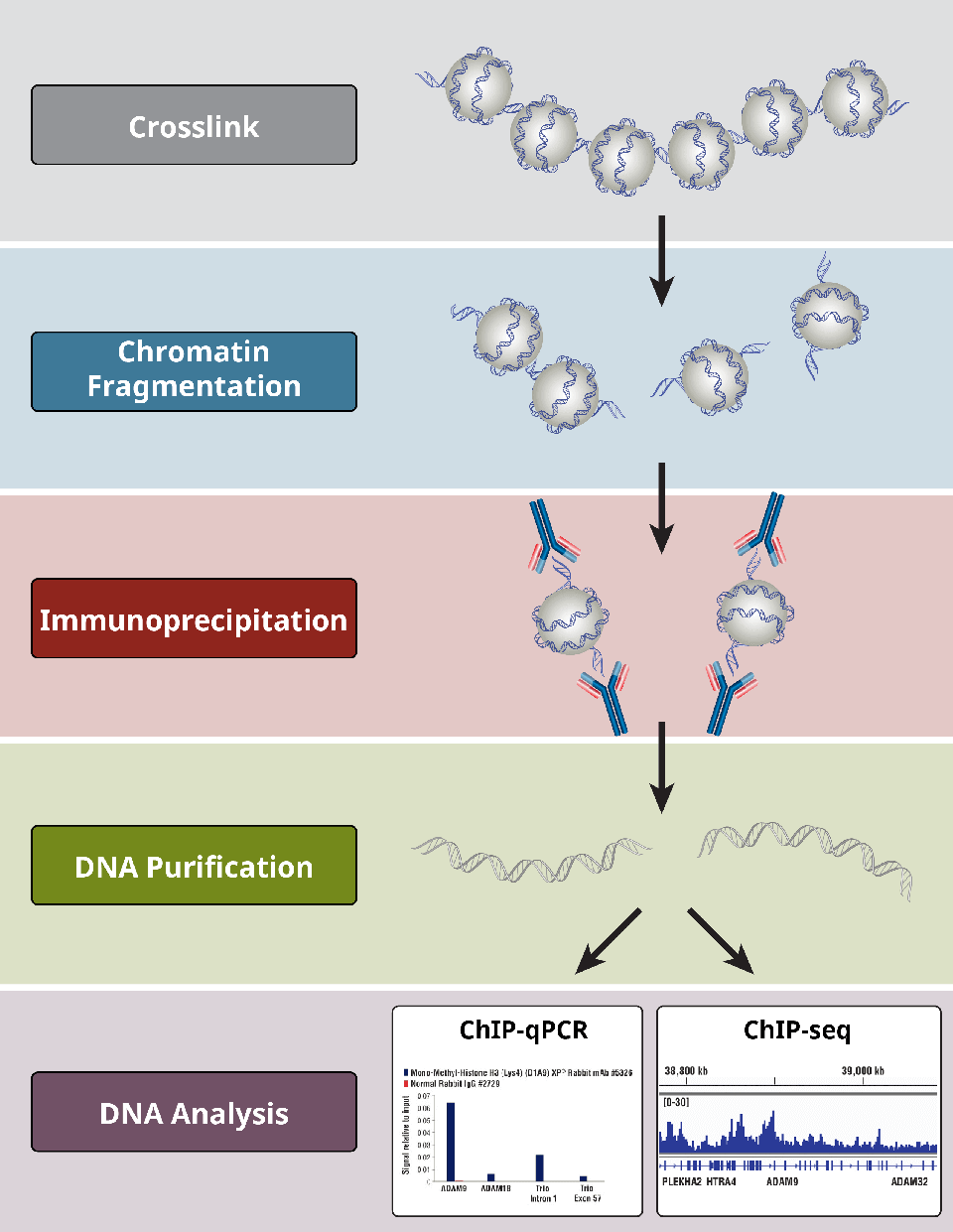
Figure 1: Flowchart of Chromatin Immunoprecipitation (ChIP) assay. First, fix the protein-DNA interaction via formaldehyde cross-linking, then fragment chromatin by sonication, capture the target protein-DNA complex with specific antibodies, obtain enriched DNA fragments after washing and purification, and finally analyze by qPCR or sequencing.
Why is ChIP the "Gold Standard" for Epigenetic Research?
When exploring gene regulatory mechanisms, researchers often face a fundamental challenge: protein-DNA interactions are dynamic and transient. In living cells, the binding of transcription factors to chromatin may last only a few minutes, and traditional chromatin isolation techniques are difficult to capture these brief interactions.
The exquisite feature of ChIP technology lies in the strategy of "lock first, then separate":
1 In Situ Fixation: Cross-linking agents such as formaldehyde penetrate cell membranes and covalently link proteins to DNA in living cells. This step is equivalent to taking a "snapshot" of the cell, instantly freezing all ongoing interactions.
2 Selective Enrichment: "Fish out" the target protein with specific antibodies, and only the DNA fragments bound to the target protein will be co-precipitated, achieving precise enrichment of specific sequences from millions of sites across the genome.
3 De-crosslinking & Purification: Reverse formaldehyde cross-linking by heating or high-salt treatment to release pure DNA, ready for subsequent analysis.
Compared with other techniques such as EMSA (Electrophoretic Mobility Shift Assay) or yeast one-hybrid, the greatest advantage of ChIP is in vivo authenticity —— it reflects the real interaction between proteins and chromatin under physiological conditions, rather than an artificially reconstructed environment in vitro.
Which Experimental Scenarios Must Use ChIP Kits?
Transcription Factor Binding Site Identification
This is the most classic application scenario of ChIP. When you study a key transcription factor in a signaling pathway, ChIP can help you identify the target genes it directly regulates. For example, in the study of inflammatory responses, ChIP assays for NF-κB can precisely locate its binding sites in the promoter regions of genes such as IL-6 and TNF-α, thus verifying its regulatory mechanism.
Histone Modification Mapping
Epigenetic studies have found that chemical modifications of histones (such as H3K4me3, H3K27ac, etc.) are closely related to gene transcriptional activity. Using antibodies against specific modifications for ChIP, genome-wide or specific gene region histone modification maps can be drawn to reveal the "active state" of chromatin.
DNA Methylation and Chromatin Remodeling Research
Although ChIP does not directly detect DNA methylation, combined with antibodies against methylated DNA-binding proteins (such as MeCP2), it can indirectly study how DNA methylation affects protein-chromatin interactions. Similarly, research on chromatin remodeling complexes (such as the SWI/SNF family) is inseparable from ChIP technology.
Regulatory Mechanism Exploration of Long Non-coding RNAs (lncRNAs)
Recent studies have found that many lncRNAs exert regulatory functions by recruiting chromatin-modifying complexes. The combined strategy of ChIP-RNA or ChIP coupled with RNA Immunoprecipitation (RIP) can reveal the three-dimensional interaction network of lncRNA-protein-DNA.
Drug Mechanism of Action Research
In the development of drugs targeting transcriptional regulation, ChIP is an important tool for evaluating drug efficacy. For example, after treatment with histone deacetylase inhibitors (HDACi), the drug effect can be directly verified at the molecular level by detecting changes in histone acetylation levels and transcription factor binding.
How to Ensure the Success Rate of ChIP Experiments?
Optimization of Cross-linking Conditions
Cross-linking is the first step determining the success or failure of ChIP. Too low formaldehyde concentration leads to incomplete cross-linking, and protein-DNA complexes dissociate in subsequent steps; too high concentration masks antigenic epitopes, making them unrecognizable by antibodies. It is generally recommended to use 1% formaldehyde for cross-linking at room temperature for 10-15 minutes, and the time can be appropriately extended for proteins that are difficult to cross-link.
The Art of Sonication
Sonication is a key step for chromatin fragmentation. Overly large fragments (>1500 bp) reduce resolution and fail to precisely locate binding sites; overly small fragments (<100 bp) may destroy antigenic epitopes. Optimizing sonication conditions requires repeated exploration, and agarose gel electrophoresis is used to confirm that the fragment size distribution is between 200-1000 bp.
Pitfalls of Antibody Selection
Not all antibodies are suitable for ChIP. Many commercial antibodies are optimized for Western blot or immunofluorescence, recognizing linear epitopes after denaturation. ChIP requires antibodies that can recognize native conformation chromatin-binding proteins, so ChIP-validated antibodies must be selected, or specificity verified by peptide competition experiments.
Scientificity of Control Setup
A rigorous ChIP experiment must include:
- Positive Control: Use antibodies against RNA polymerase II or histone H3, which are stably enriched in the promoter regions of active genes
- Negative Control: Use normal IgG or samples without primary antibody to evaluate non-specific background
- Input Control: Take a small amount of unimmunoprecipitated chromatin as a control for subsequent quantitative calculation of enrichment folds
From ChIP to ChIP-seq: The Evolution of Technology
With the development of high-throughput sequencing technology, ChIP is no longer limited to the detection of single or a few loci. ChIP-seq (Chromatin Immunoprecipitation Sequencing) combines ChIP with genome-wide sequencing, enabling the mapping of target protein binding profiles across the entire genome in a single experiment.
For ChIP-seq, the quality of DNA purification provided by the kit is particularly critical. Sequencing library construction has strict requirements on the size distribution and purity of DNA fragments, and any residual protein or salt may affect the efficiency of subsequent adapter ligation and PCR amplification. Therefore, choosing a kit with high-quality purification columns is the foundation for successful ChIP-seq.
Conclusion
Chromatin Immunoprecipitation kits provide researchers with a key to unlock the black box of gene regulation. From the precise localization of transcription factors to the panoramic scanning of epigenetic modifications, from the in-depth analysis of single genes to genome-wide mapping, ChIP technology has always been the most reliable method for verifying protein-DNA interactions. Today, with the vigorous development of epigenetics, mastering ChIP technology is not only an improvement in experimental skills, but also a necessary path to deeply understand the mechanism of gene expression regulation.
Absin Immunoprecipitation Kit Recommendation
Contact Absin
Absin provides antibodies, proteins, ELISA kits, cell culture, detection kits, and other research reagents. If you have any product needs, please contact us.
| Absin Bioscience Inc. worldwide@absin.cn |
 Follow us on Facebook: Absin Bio Follow us on Facebook: Absin Bio |
