- Cart 0
- English
Single-Molecule RNA Fluorescence In Situ Hybridization (GeneScope): A Cutting-Edge Technology for Precisely Analyzing the Spatiotemporal Expression of RNA
May 08, 2026
Clicks:73
In life science research, precisely localizing and quantifying RNA expression levels in tissues and cells is critical for understanding gene regulation, cellular functions, and disease pathogenesis. Single-molecule RNA fluorescence in situ hybridization (amsFISH, also commonly known as smFISH) has become a core tool in this field due to its ultra-high sensitivity and single-molecule resolution. The GeneScope Multicolor Single-Molecule RNA FISH Kit launched by Absin (absin) provides a highly efficient solution for this advanced technology.
1. Core Principle of amsFISH Technology
The core principle of amsFISH is the specific hybridization of multiple pairs of short DNA probes with target RNA molecules, followed by signal amplification and fluorescence imaging to enable visualization and quantitative analysis of single RNA molecules.
1. Probe Design and Hybridization: 5-10 pairs (or more) of short, labeled DNA probes are designed against the target RNA sequence. These probes bind simultaneously to different sites of the same RNA molecule, forming stable hybrid complexes.
2. Signal Amplification: Unlike traditional FISH, amsFISH employs rolling circle amplification (RCA) and other techniques to cascade-amplify the signal of each hybridized probe. This allows even low-abundance RNA molecules to generate sufficiently strong fluorescent signals for clear microscopic detection.
3. Single-Molecule Imaging and Quantification: Each fluorescent spot represents one RNA molecule. High-resolution fluorescence microscopy and dedicated analysis software enable accurate counting and spatial localization of RNA molecules in cells or tissue sections, achieving true single-cell and single-molecule quantification.
2. How GeneScope Empowers amsFISH Experiments
Absin's GeneScope Multicolor Single-Molecule RNA FISH Kit streamlines and standardizes the complex amsFISH workflow, greatly lowering experimental barriers while ensuring reliable results.
 Schematic diagram of double-stranded DNA probe circularization and amplification for signal enhancement
Schematic diagram of double-stranded DNA probe circularization and amplification for signal enhancement
1. Technical Advantages: Integration of Precision and Efficiency
GeneScope fully embodies and optimizes the technical strengths of amsFISH:
- High Sensitivity: Stably detects low-abundance target RNAs, solving the "invisibility" issue of low-expression genes with traditional methods.
- High Specificity: Exclusive probe design effectively inhibits non-specific binding, ensuring signals truly reflect target RNA localization.
- Single-Cell Quantification: Precise expression analysis at the single-cell level reveals heterogeneity within cell populations.
- Morphology Preservation: Maximally retains intact tissue and cellular morphology during RNA detection, facilitating spatial distribution analysis of gene expression.
2. Workflow: One-Stop Service from Customization to Analysis
GeneScope features a clear and efficient workflow with only 5 core steps:
- Custom Probe Design: Free probe design and synthesis based on your provided species, gene names, and sequence information.
- Sample Preprocessing: Fixation, permeabilization, and other pretreatment of tissue sections or cell samples.
- Probe Hybridization and Rolling Circle Amplification: Incubate probes with samples, followed by rolling circle amplification for signal enhancement.
- In Situ Imaging: Image acquisition using fluorescence microscopy.
- Image Analysis: Counting and analysis of fluorescent spots via specialized software.
Notably, the entire workflow only requires conventional instruments such as constant-temperature incubators, vortex mixers, and palm centrifuges—no expensive hybridization ovens are needed, significantly reducing laboratory equipment investment.
3. Product Ordering and Supporting Kits (Details in the poster below)
GeneScope offers flexible ordering plans and comprehensive supporting kits to meet diverse experimental needs:
- Flexible Customization: Simultaneously detect 1-4 RNA types (including mRNA, lncRNA, circRNA, etc.) in one experiment; probe customization takes only 7 working days.
- Supporting Kits: Absin provides complete detection kits for 1 to 4 targets (Cat. Nos. abs60728-abs60731), containing all essential components for sample preprocessing, reaction systems, target repair, and complimentary positive/negative control probes—ready to use out of the box.
3. Application Scenarios: From Developmental Biology to Cancer Research
amsFISH technology, especially high-efficiency tools like GeneScope, has excelled in multiple cutting-edge research fields:
- Developmental Biology: In developmental biology research, single-molecule RNA FISH is a key tool for decoding the spatiotemporal regulation of embryonic development, enabling precise RNA localization and cellular heterogeneity analysis in intact tissues.
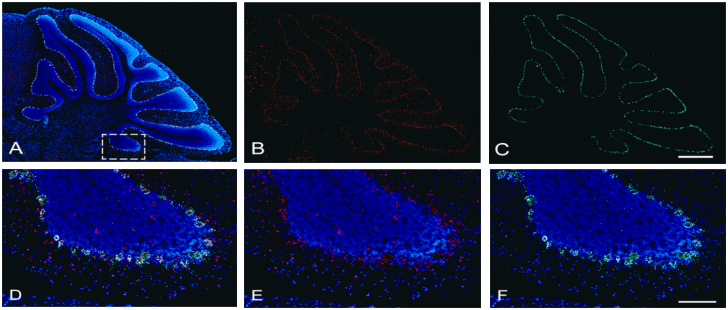
- Neuroscience: For example, the diagram below shows localization of two genes in frozen brain tissue sections: Gad1 (red), glutamate dehydrogenase 1, highly expressed in the molecular layer and some granular layer cells; Calb1 (green), calbindin 1, mainly present in the Purkinje cell layer, with low expression in the molecular layer from Purkinje cell axons.
- Cancer Research: Analyzes gene expression in different cell subsets within the tumor microenvironment, revealing tumor heterogeneity and drug resistance mechanisms.
- Virology: Detects and localizes viral RNA replication sites and transmission pathways in host cells.
Contact Absin
Absin provides antibodies, proteins, ELISA kits, cell culture, detection kits, and other research reagents. If you have any product needs, please contact us.
| Absin Bioscience Inc. worldwide@absin.cn |
 Follow us on Facebook: Absin Bio Follow us on Facebook: Absin Bio |
