- Cart 0
- English
How does the rapid endonuclease XhoI enable more efficient and precise DNA digestion?
May 08, 2026
Clicks:73
Molecular cloning is a core technology in modern life science research, and restriction endonucleases are the "molecular scissors" for molecular cloning. Traditional restriction endonuclease reactions usually take 1-2 hours or even longer, and different enzymes require different reaction buffers, making the operation cumbersome and time-consuming. Through genetic engineering modification, the fast restriction endonuclease series achieves efficient DNA cleavage within 5-15 minutes, greatly improving experimental efficiency. This article will deeply explore the characteristics, recognition mechanism and application of fast restriction endonuclease XhoI in molecular cloning.
What is Fast Restriction Endonuclease XhoI?
Fast restriction endonuclease XhoI is a genetically engineered restriction enzyme that can precisely complete DNA cleavage within 5-15 minutes. This enzyme recognizes specific DNA sequences and cleaves double-stranded DNA at defined positions to generate DNA fragments suitable for subsequent cloning operations.
Recognition site of XhoI:
3'...GAGCT↑C...5'
The arrows indicate the cleavage sites, generating 5' overhang sticky ends (5'-TCGA-3'). Such ends can be ligated with compatible ends produced by enzymes such as SalI and XhoI.
Isoschizomers of XhoI include PaeR7I, TliI, BssHI, Sfr274I, SlaI, StrI, etc. These enzymes recognize the same DNA sequence and generate identical ends, and can be used interchangeably in experimental design.
The enzyme activity is 20 U/μL. The product usually contains XhoI enzyme solution (500μL), 10X Cut Buffer (31mL) and 10X Cut Color Buffer (31mL), stored at -20°C with a validity period of 2 years.
What are the Technical Advantages of Fast Restriction Endonucleases?
Rapid reaction is the most prominent feature of fast restriction endonucleases. Digestion can be completed within 5-15 minutes, greatly shortening the experimental time compared with 1-2 hours of traditional enzymes. Plasmid DNA is usually digested in 15 minutes, PCR products in 15-30 minutes, and genomic DNA in 30-60 minutes.
Universal buffer system simplifies the operation process. All fast restriction endonucleases share one digestion buffer, eliminating the need to change buffers for different enzymes and greatly simplifying the preparation of digestion reaction systems. In addition, supporting dephosphorylation and ligation reagents exhibit 100% activity in the supporting buffer, enabling "one-tube" digestion-modification-ligation reactions.
Excellent enzyme activity redundancy ensures complete digestion. The enzyme preparation has sufficient activity reserve to easily handle excessive substrates or difficult template digestion, guaranteeing cleavage efficiency.
High purity reduces non-specific cleavage. Through strict quality control, no non-specific substrate degradation caused by other nuclease contamination or star activity is detected.
In Which Experiments Can XhoI Be Used?
Plasmid DNA digestion is the most basic application of XhoI. In vector construction, XhoI is used to cleave plasmid vectors to generate specific sticky ends for inserting target gene fragments. Since XhoI usually has no or rare restriction sites in common cloning vectors (such as pUC series, pBR322, etc.), it is an ideal choice for multiple cloning sites.
PCR product digestion is used for cloning amplified gene fragments. After purification, PCR products are cleaved by XhoI to generate ends compatible with vectors, facilitating directional cloning. Note that unpurified PCR products have a certain ionic strength, and the volume of 10X Cut Buffer can be appropriately reduced; however, since DNA polymerase also has exonuclease activity that affects digestion products, purification of PCR products before digestion is recommended for subsequent cloning operations.
Genomic DNA digestion is used in experiments such as Southern Blot and genomic library construction. Genomic DNA has a complex structure, and the digestion time needs to be appropriately extended to 30-60 minutes.
Double or multiple digestion is a routine operation for vector construction. XhoI can perform double or multiple digestion with other fast restriction endonucleases in the same reaction system, and the total volume of all fast restriction endonucleases shall not exceed 1/10 of the total reaction system. If the optimal reaction temperatures of the enzymes are different, perform digestion with the enzyme having the lower optimal temperature first, then add the enzyme with the higher optimal temperature.
Digestion-ligation-redigestion verification is used to confirm digestion and ligation efficiency. Digest the substrate with XhoI, recover the digested products, religate with T4 DNA ligase, and re-digest the ligated products with the same restriction endonuclease to verify the correctness of the ligation products.
In blue-white screening, XhoI is used to cleave vectors containing the lacZα gene. Correctly ligated products produce blue colonies, while incorrectly ligated products (i.e., incomplete DNA ends) yield white colonies, with the white colony ratio less than 1%.
How to Use Fast Restriction Endonuclease XhoI Correctly?
Reaction System Preparation (Taking Plasmid DNA as an Example):
| Component | Volume |
|---|---|
| ddH₂O | 15 μL |
| 10X Cut Buffer or 10X Cut Color Buffer | 2 μL |
| Substrate DNA | 2 μL (up to 1 μg) |
| XhoI | 1 μL |
| Total | 20 μL |
Operating Steps:
- Prepare the reaction system on ice in sequence
- Mix gently by pipetting or flicking the tube wall (do not vortex), then centrifuge briefly to collect liquid droplets on the tube wall
- Incubate at 37°C for 15 minutes (plasmid), or 15-30 minutes (PCR product), or 30-60 minutes (genomic DNA)
- Inactivate the enzyme by incubating at 80°C for 20 minutes to stop the reaction (optional)
Double Digestion Operation:
The dosage of each fast restriction endonuclease is 1μL, and the reaction system can be appropriately expanded as needed. The total volume of all fast restriction endonucleases shall not exceed 1/10 of the total reaction system. If the optimal temperatures of the enzymes are different, perform digestion with the enzyme having the lower optimal temperature first, then add the enzyme with the higher optimal temperature.
What are the Key Precautions for Use?
- Methylation modification effect. XhoI is sensitive to CpG methylation, and cleavage is inhibited when sequences are completely overlapped. If DNA is derived from higher eukaryotes such as mammals, its CpG sites are usually methylated, which may affect the digestion efficiency of XhoI. Dam and Dcm methylation have no effect on XhoI.
- Reaction temperature control. 37°C is the optimal reaction temperature for XhoI. If the total reaction system is larger than 20μL, the incubation time should be appropriately increased, and water bath, metal bath or sand bath should be used as much as possible to ensure uniform temperature.
- Enzyme inactivation method. The enzyme can be inactivated by incubating at 80°C for 20 minutes. The reaction can also be terminated by phenol-chloroform extraction or agarose gel electrophoresis recovery.
- Avoid repeated freeze-thaw cycles. The enzyme solution should be stored at -20°C, and repeated freeze-thaw cycles should be avoided to prevent activity reduction. Aliquoting into small volumes is recommended for storage.
- Sterile operation. Prepare the reaction system under sterile conditions to prevent DNA contamination.
- Quality control verification. Functional activity assay shows that 1μL XhoI can completely digest 1μg λDNA within 15 minutes in a 20μL reaction system. Ultra-long incubation assay shows no other nuclease contamination or star activity detected after 3-hour incubation.
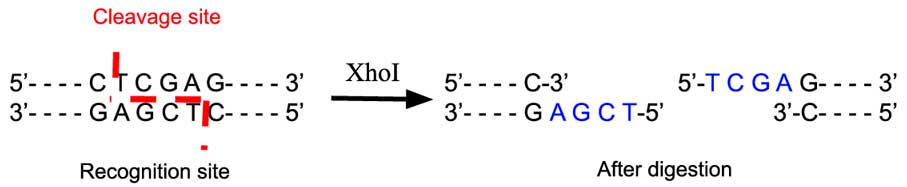
Schematic diagram of recognition and cleavage by XhoI restriction endonuclease
Conclusion
Fast restriction endonuclease XhoI realizes rapid, standardized and simplified DNA digestion reactions through genetic engineering modification. The 5-15 minute rapid reaction, universal buffer system and excellent enzyme activity redundancy make it an efficient tool for molecular cloning experiments. From plasmid construction to genomic analysis, from single digestion to combined multiple digestion, XhoI plays an important role in genetic engineering, protein expression, vector construction and other fields. Correct mastery of digestion reaction conditions, attention to methylation modification effects and operational details will ensure digestion efficiency and cloning success rate. With the development of synthetic biology and gene editing technology, restriction endonucleases, as basic tools for molecular cloning, will continue to show their irreplaceable value in life science research.
Absin Fast Restriction Endonuclease XhoI Recommendation:
| Cat. No. | Product Name | Size |
|---|---|---|
| abs60243 | Fast Restriction Endonuclease XhoI | 500T |
Contact Absin
Absin provides antibodies, proteins, ELISA kits, cell culture, detection kits, and other research reagents. If you have any product needs, please contact us.
| Absin Bioscience Inc. worldwide@absin.cn |
 Follow us on Facebook: Absin Bio Follow us on Facebook: Absin Bio |
