- Cart 0
- English
How do Type IIS restriction enzymes enable seamless and precise assembly of DNA fragments?
April 30, 2026
Clicks:74
Throughout the continuous evolution of molecular cloning technology, restriction endonucleases have always been the core tool for DNA manipulation. Although traditional Type II restriction enzymes (such as EcoRI, HindIII) are widely used, their overlapping recognition and cleavage sites often result in residual restriction site "scars" in cloned products, limiting the possibility of seamless multi-fragment assembly. As a typical Type IIS restriction enzyme, BsaI enables precise, efficient and scarless assembly of DNA fragments through the spatial separation of recognition and cleavage sites combined with fast digestion technology, providing a powerful technical platform for synthetic biology, gene editing and the construction of complex genetic circuits.
Why can Type IIS restriction enzymes achieve seamless cloning?
Restriction enzymes are classified based on their recognition sequence characteristics, cleavage site position and cofactor requirements. The uniqueness of Type IIS restriction enzymes lies in their non-palindromic recognition sequences and cleavage properties outside the recognition site:
Spatial separation of recognition and cleavage:
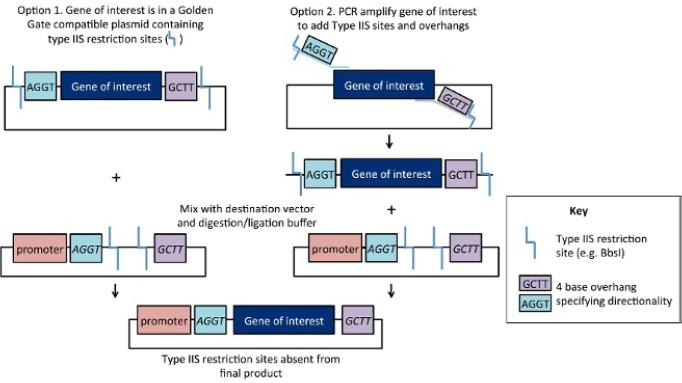
BsaI recognizes the 5'-GGTCTC-3' sequence, but the cleavage site is located downstream of the recognition sequence: it cleaves the top strand 1 base after the recognition site (N1) and the bottom strand 5 bases after (N5), generating 4-base 3' overhang sticky ends. This "recognize without cutting, cut without leaving scars" feature ensures that the recognition sequence is completely removed after digestion and does not appear in the final assembled product.
Customizable sticky ends:
Since the cleavage site is independent of the recognition sequence, the 4-base overhang sequence can be arbitrarily designed (256 possibilities in total), guiding the ordered assembly of multiple fragments through complementary pairing. This flexibility allows a single enzyme to complete directional multi-fragment cloning without considering internal restriction sites of the fragments.
Kinetic advantages of cyclic reactions:
Golden Gate assembly alternates between digestion and ligation in the same reaction system: DNA is cleaved at the optimal digestion temperature (37°C), and fragments are annealed and ligated at the optimal ligation temperature (16°C or room temperature). Since correctly ligated products no longer contain recognition sites and cannot be recleaved, correct products accumulate continuously as the reaction proceeds, with efficiency exceeding 90%.
How does fast digestion technology improve cloning efficiency?
High enzyme activity and buffer compatibility:
Traditional restriction enzyme reactions usually take 1-2 hours, while fast endonucleases achieve efficient digestion in 5-15 minutes through protein engineering modification. Genetically engineered BsaI maintains high activity in various buffers, including dedicated Cut Buffer, T4 DNA Ligase Buffer, and NEB CutSmart® Buffer, with 100% activity. This buffer compatibility allows digestion-ligation reactions to proceed in the same system without intermediate buffer replacement or DNA purification.
Reduced star activity:
Star activity refers to non-specific cleavage by restriction enzymes under suboptimal conditions, a major interference in cloning experiments. High-fidelity (HF) version of BsaI reduces star activity through amino acid substitution and improves cleavage specificity.
Temperature flexibility and stability:
The optimal reaction temperature of BsaI is 37°C, but it maintains activity over a wide temperature range. After reaction completion, the enzyme can be inactivated by incubation at 80°C for 20 minutes to terminate the reaction.
Which experimental scenarios best reflect its technical advantages?
Golden Gate seamless cloning assembly
Golden Gate cloning is the most classic application of BsaI. By introducing BsaI recognition sequences and protective bases at the 5' end of primers, PCR products can be digested to generate specific sticky ends, achieving seamless ligation with similarly treated vectors under the action of T4 DNA Ligase. This technology can assemble 2-20+ fragments and has been successfully applied to complex assembly of 52 fragments.
TALEN and CRISPR vector construction
Construction of transcription activator-like effector nucleases (TALEN) and CRISPR systems requires precise assembly of multiple repeat sequence modules. The Golden Gate method uses BsaI to generate specific ends, efficiently assembling TALE repeat units to construct nucleases targeting specific DNA sequences.
Synthetic biology standard part assembly
Standardized assembly systems developed based on the Golden Gate principle, such as MoClo, Golden Braid, and Loop assembly, adopt Type IIS enzymes like BsaI to establish universal ligation syntax, enabling modular assembly of biological elements (promoters, coding sequences, terminators, etc.) like "building blocks". This standardization greatly promotes the sharing and reuse of biological elements.
Library construction and screening
BsaI can be used to construct diverse DNA libraries. By introducing degenerate sequences into primers, diverse sticky ends are generated after digestion, forming libraries containing multiple combinations after ligation for protein engineering or regulatory element screening.
How to optimize digestion and assembly experiments?
Primer design and protective bases
When introducing BsaI recognition sequences at both ends of the target fragment, 3-6 protective bases should be added at the 5' end to ensure stable enzyme binding and efficient cleavage. Common protective base sequence: GGTCTCN↓ (N = any base, ↓ = cleavage site).
Reaction system preparation
Plasmid DNA digestion (20μL system):
- ddH₂O: 15μL
- 10X Cut Buffer: 2μL
- Plasmid DNA (≤1μg): 2μL
- BsaI (20U/μL): 1μL
- Incubate at 37°C for 15 minutes
PCR product digestion (30μL system):
- ddH₂O: 16μL
- 10X Cut Buffer: 3μL
- PCR product (~0.2μg): 10μL
- BsaI: 1μL
- Incubate at 37°C for 15-30 minutes
Genomic DNA digestion (50μL system):
- ddH₂O: 30μL
- 10X Cut Buffer: 5μL
- Genomic DNA (5μg): 10μL
- BsaI: 5μL
- Incubate at 37°C for 30-60 minutes
Notes on multi-enzyme digestion reactions
For simultaneous double or multi-enzyme digestion:
- Use 1μL of each enzyme, total enzyme volume ≤ 1/10 of the reaction system
- If enzymes have different optimal temperatures, start with the low-temperature enzyme first, then add the high-temperature enzyme for continued reaction
- Ensure all enzymes are active in the selected buffer (BsaI has 100% activity in CutSmart® Buffer)
Methylation sensitivity
BsaI is sensitive to certain methylation modifications:
- Dam methylation: No effect
- Dcm methylation: Cleavage blocked when sequences fully overlap
- CpG methylation: Cleavage blocked when sequences fully overlap
When using mammalian genomic DNA or methylated plasmids, confirm whether methylation modifications exist at restriction sites; if necessary, perform demethylation treatment or select alternative enzymes.
Type IIS restriction enzyme BsaI recognizes the non-palindromic sequence GGTCTC and cleaves outside the recognition site to generate 4-base specific sticky ends. Due to the separation of cleavage and recognition sites, the recognition sequence is completely removed after digestion, and multiple fragments can be directionally assembled through complementary sticky ends to form a seamless ligation product
Considerations for technology selection
When selecting BsaI for cloning experiments, comprehensively evaluate the following factors:
Fragment number:
Single-fragment cloning is simple and efficient; multi-fragment assembly requires careful sticky end design to avoid self-ligation and mispairing.
Sequence content:
Check for internal BsaI recognition sites in inserts and vectors; if present, remove them via silent mutation (domestication).
Downstream applications:
For in vitro transcription, select dedicated vectors containing T7/SP6 promoters; for protein expression, ensure correct open reading frame fusion.
Time efficiency:
Fast digestion version (5-15 minutes) is suitable for high-throughput experiments; standard version is ideal for routine laboratory use.
Conclusion
Fast endonuclease BsaI represents the evolution of restriction enzyme technology from "molecular scissors" to "molecular scalpel". Through the unique cleavage mechanism of Type IIS type and protein engineering optimization, BsaI achieves rapid, precise and scarless assembly of DNA fragments, providing an efficient platform for modern molecular cloning. From single-gene cloning to multi-fragment synthetic biology circuits, from basic research to applied development, BsaI and its Golden Gate assembly technology are driving genetic engineering toward higher throughput and precision. Mastering its principles and operational key points helps researchers achieve twice the result with half the effort in complex DNA assembly tasks.
Absin Fast Endonuclease BsaI Recommendation:
| Cat. No. | Product Name | Size |
|---|---|---|
| abs60206 | Fast Endonuclease BsaI | 50T |
Contact Absin
Absin provides antibodies, proteins, ELISA kits, cell culture, detection kits, and other research reagents. If you have any product needs, please contact us.
| Absin Bioscience Inc. worldwide@absin.cn |
 Follow us on Facebook: Absin Bio Follow us on Facebook: Absin Bio |