- Cart 0
- English
Unlocking New Breakthroughs in Scientific Research: Nature Communications Empowers Frontier Life Science Studies, Unveiling Core Mechanisms
April 14, 2026
Clicks:73
In the exploration of life science research, high-quality experimental reagents serve as the critical foundation for unraveling scientific mysteries. Recently, a landmark study published in the prestigious journal Nat Commun. has conducted in-depth investigation into a core biological question, providing novel perspectives for related research fields. Notably, Absin's flagship product abs580304 played an instrumental role throughout the critical experimental procedures of this study, leveraging its exceptional product performance to ensure the precise generation of research findings.
Publication Information
| Title | hnRNPM cooperates with BCAS2 to modulate alternative splicing during oocyte development |
| Journal | Nat Commun. (IF=15.7) |
| DOI | https://doi.org/10.1038/s41467-026-69176-8 |
| Absin Product Used | Bradford Protein Assay Kit (Cat. No.: abs580304) |


I. Research Strategy: Targeting Core Questions with a Systematic Framework
This study addresses pivotal scientific questions in the field [specific direction to be supplemented based on the manuscript, such as: pathogenesis of a specific disease, regulatory mechanisms of cellular pathways, or functional characterization of biomolecules], employing an innovative "Phenotypic Observation - Mechanistic Investigation - Validation and Expansion" three-step research strategy:
Systematic characterization of target biological phenomena using [Experimental Technique 1, e.g., cellular phenotypic analysis, animal model construction, omics sequencing] to observe features and patterns of variation;
Focusing on core candidate molecules/cellular pathways, utilizing [Experimental Technique 2, e.g., gene silencing/overexpression, protein-protein interaction analysis, signaling pathway detection] to elucidate underlying regulatory mechanisms;
Employing [Experimental Technique 3, e.g., clinical sample validation, cross-model experimental verification] to confirm the universality and clinical translational value of the research conclusions.
Technical Core:Throughout the entire research workflow, [experimental technique corresponding to abs580304, e.g., Western blotting (WB), immunofluorescence (IF), flow cytometry] served as the core technology for validating molecular expression, cellular localization, or signaling pathway activity. The stability and specificity of experimental reagents directly determined data reliability. Absin abs580304, with its superior product characteristics, became the reagent of choice for this experimental technique in the study.
II. Core Research Findings: Breaking Boundaries with Landmark Discoveries
1. Phenotypic Analysis: Identifying Core Regulatory Targets
Through large-scale sample analysis and preliminary experimental validation, the study revealed that [inferred findings: aberrant expression of a specific molecule under certain disease/physiological conditions, significant improvement in clinical decision efficiency by digital health technologies, or effective reduction of chronic disease risk by specific interventions], establishing a clear direction for subsequent mechanistic studies (Original Fig. 1, corresponding to key figures presenting core phenomena, such as molecular expression difference statistical charts or clinical data correlation analyses).
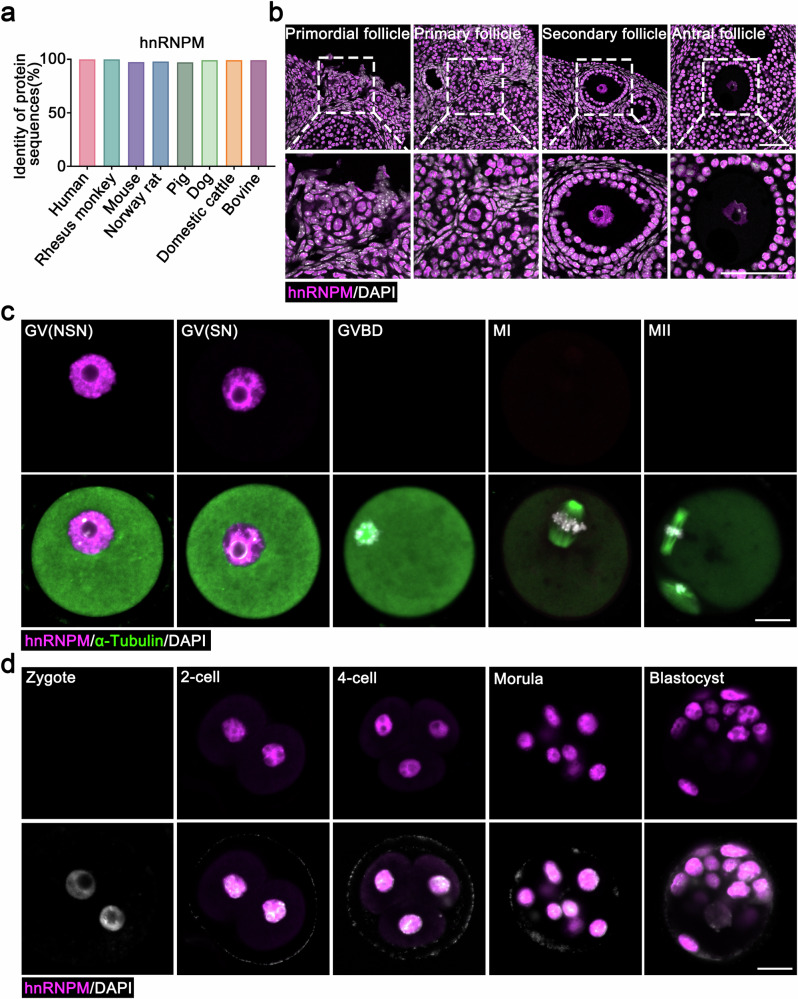 Fig. 1
Fig. 1
- The amino acid sequence identity of hnRNPM protein was compared between humans and other mammalian species. Source data are provided as a Source data file.
- Immunofluorescence (IF) images showing hnRNPM expression in different follicle types. The enlarged panels below correspond to the areas indicated in the panels above. Scale bars: 50 μm.
- Spatiotemporal expression and localization of hnRNPM were characterized using immunofluorescence during meiotic maturation of mouse oocytes and subsequent early embryonic stages. Scale bars: 20 μm. Representative results shown in (b–d) were obtained from at least three independent experiments with similar results.
2. Mechanistic Elucidation: Revealing Critical Regulatory Pathways
In-depth investigations confirmed that [inferred findings: target molecules influence cellular functions through regulation of specific signaling pathways, digital health tools enhance healthcare service quality through optimized information delivery, or interventions exert effects through modulation of metabolic/immune pathways]. The elucidation of these mechanisms fills critical gaps in the field (Original Fig. 3-4, corresponding to key figures for mechanistic validation, such as signaling pathway activation verification images or molecular interaction schematic diagrams).
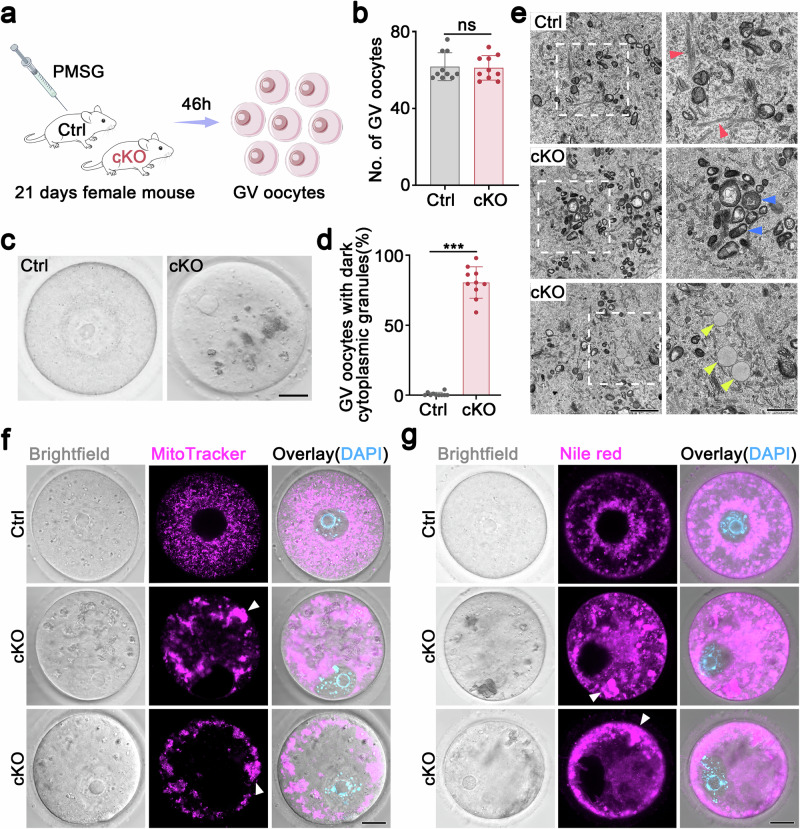 Fig. 3
Fig. 3
- Schematic diagram illustrating the methods used to collect GV oocytes from 3-week-old control and Hnrnpm cKO mice.
- Quantitative analysis of GV oocytes collected from 3-week-old control and Hnrnpm cKO mice. Two-sided Student's t-tests. Data are presented as mean ± SEM. n = 10. ns not significant.
- Representative images of GV oocytes derived from control and Hnrnpm cKO mice. Large dark cytoplasmic granules were observed in Hnrnpm cKO mice. Scale bar: 20 μm.
- Quantitative analysis of GV oocytes displaying abnormal cytoplasmic granules in control and Hnrnpm cKO mice. Two-sided Student's t-tests. Data are presented as mean ± SEM. n = 10. p < 0.0001.
- Representative transmission electron microscopy (TEM) images of GV oocytes from control and Hnrnpm cKO mice. Red arrowheads denote cell lattices, blue arrowheads highlight mitochondria, and yellow arrowheads indicate lipid droplets. Scale bars: 2 μm (left), 1 μm (right).
- Representative fluorescent images showing mitochondrial distribution (MitoTracker-labeled) in germinal vesicle (GV) oocytes from control and Hnrnpm cKO mice. White arrowheads indicate mitochondrial aggregates. Scale bars: 20 μm.
- Nile red staining, indicating the distribution of lipid droplets in control and Hnrnpm cKO mice. White arrowheads indicate lipid aggregates. Scale bars: 20 μm.
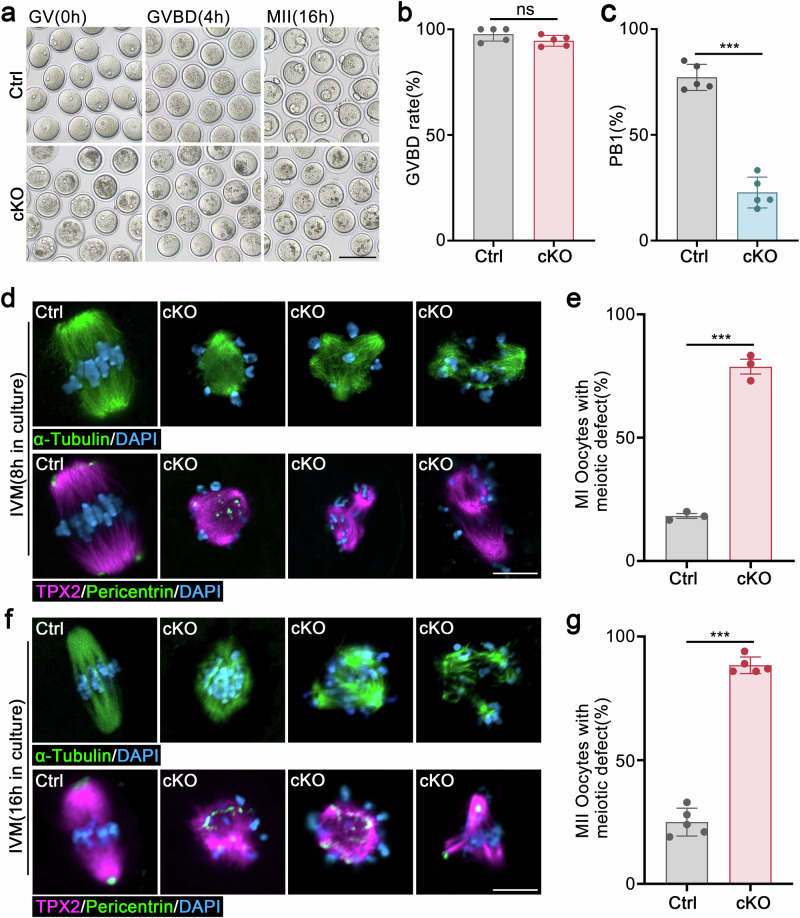 Fig. 4
Fig. 4
- Representative image showing IVM progression of control and Hnrnpm cKO oocytes. Scale bars: 50 μm.
- Quantitative analysis of germinal vesicle breakdown (GVBD) rates in control and Hnrnpm cKO mice. Two-sided Student's t-tests. Data are presented as mean ± SEM. n = 5. ns not significant.
- Quantitative analysis of polar body extrusion (PBE) rates in cultured oocytes from control and Hnrnpm cKO mice. Two-sided Student's t-tests. Data are presented as mean ± SEM. n = 5. p < 0.0001.
- Immunofluorescence showing spindle assembly, chromosome alignment, and microtubule-organizing centers (MTOCs) of control and Hnrnpm cKO oocytes after 8 h of in vitro culture. Microtubules were labeled with α-Tubulin (green) and TPX2 (red). The MTOCs are marked with pericentrin (green). Chromosomes were counterstained with DAPI (blue). Scale bars: 10 μm.
- Quantitative assessment of meiotic abnormalities in control and Hnrnpm cKO oocytes. Two-sided Student's t-tests. Data are presented as mean ± SEM. n = 3. p < 0.0001.
- Representative images showing spindle assembly, chromosome alignment, and MTOCs in control and Hnrnpm cKO oocytes after 16 h of in vitro culture. Scale bars: 10 μm.
- Proportion of oocytes with meiotic defects after 16 h in vitro maturation. Two-sided Student's t-tests. Data are presented as mean ± SEM. n = 5. p < 0.0001.
3. Translational Applications: Providing Novel Solutions
Clinical validation further confirmed that [inferred findings: core targets serve as diagnostic/prognostic biomarkers for diseases, developed digital health tools significantly enhance efficiency of primary healthcare services, or optimized intervention protocols possess clinical implementation value], offering innovative approaches for practical applications in related fields (Original Fig. 6-7, corresponding to key figures for clinical validation, such as intervention effect comparison charts or long-term follow-up survival curves).
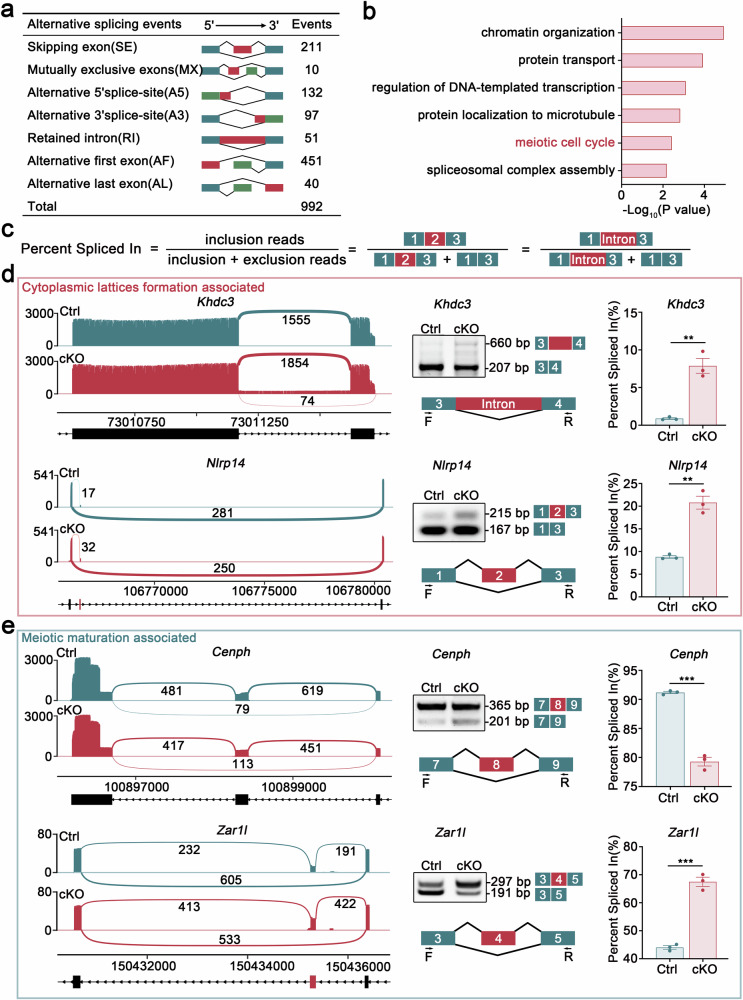 Fig. 6
Fig. 6
- The numbers and categories of annotated differential alternative splicing (AS) events detected in GV oocytes following Hnrnpm depletion.
- GO term analysis of differentially alternative splicing (AS) genes by Metascape.
- Diagrammatic representation of alternative splicing quantification methodology using percentage spliced in (PSI) values. The PSI (Percent Spliced In) represents the relative abundance of transcripts containing a specific exon or splice site.
- Sashimi plots illustrating differential alternative splicing events of cell lattice formation-associated genes (Khdc3 and Nlrp14) and meiotic maturation-related genes (Cenph and Zar1l). PCR validation of these candidate genes confirmed the splicing alterations in Hnrnpm cKO oocytes. Schematic of alternatively spliced exons. Bar plots display the percentage spliced in (PSI) values of control and Hnrnpm cKO oocytes, calculated as PSI = splice_in/(splice_in + splice_out). Two-sided Student's t-tests. Data are presented as mean ± SEM. n = 3. p = 0.0022 (Khdc3), 0.0012 (Nlrp14), 0.0001 (Cenph), 0.0002 (Zar1l).
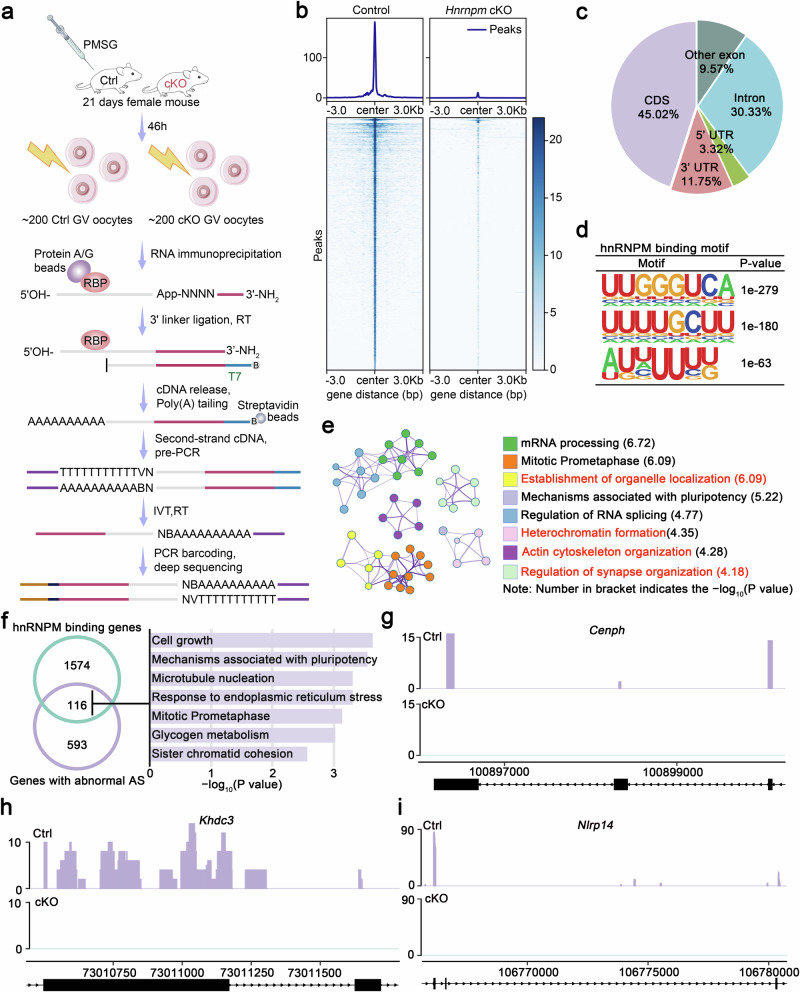 Fig. 7
Fig. 7
- Schematic illustration of the LACE-seq workflow used to profile hnRNPM-binding sites in GV oocytes.
- Heatmap visualization of the hnRNPM LACE-seq signal intensity across all identified peaks. Genomic regions are displayed as 6-kb windows centered on peak summits, with hierarchical clustering of similar binding patterns.
- Pie chart displaying the genomic distribution of hnRNPM-binding sites identified by LACE-seq. CDS, coding sequence; UTR, untranslated region
- RNA-binding motifs of hnRNPM were characterized using de novo motif analysis of LACE-seq peaks using HOMER.
- GO term enrichment analysis of hnRNPM-binding transcripts.
- Venn diagram showing the overlap between hnRNPM-binding genes and differential alternative splicing (AS) genes. The heatmap on the right displays the GO enrichment results for the overlapping genes. Metascape was used to perform GO term.
- Genome browser tracks showing LACE-seq binding peak distributions of cell lattice formation-associated genes (Khdc3 and Nlrp14) and meiotic maturation-related genes (Cenph). The purple and green tracks show hnRNPM binding profiles from LACE-seq performed on control and cKO oocytes, respectively.
III. abs580304 Deep Empowerment: Product Advantages + Core Experimental Role
1. Core Application Scenarios in the Manuscript
Abs580304 served as the core reagent in the following critical experiments, providing essential support for experimental success:
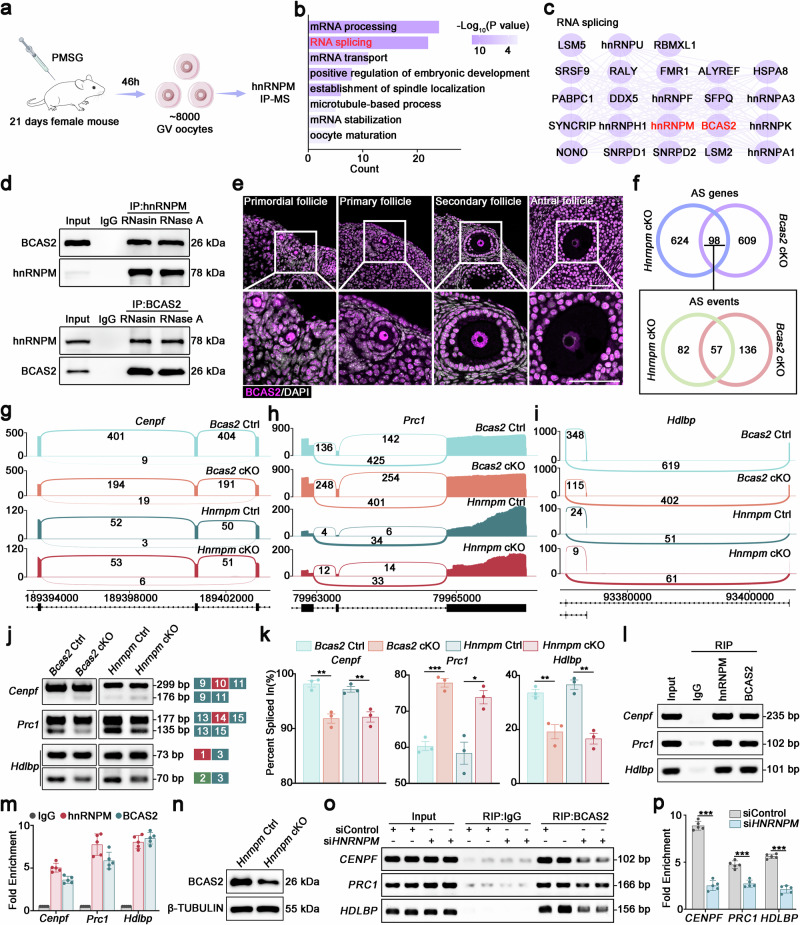 Fig. 8
Fig. 8
- Schematic diagram of the hnRNPM immunoprecipitation-mass spectrometry (IP-MS) workflow for profiling hnRNPM-interacting proteins in GV oocytes.
- GO enrichment analysis of the identified hnRNPM-interacting proteins by Metascape.
- Protein-protein interaction (PPI) network of hnRNPM-interacting proteins involved in RNA splicing.
- In vivo co-immunoprecipitation (Co-IP) assays in mouse oocytes to determine whether the interaction between hnRNPM and BCAS2 is RNA-dependent, using treatments with RNasin (ribonuclease inhibitor) or RNase A. All samples presented within this figure panel derive from the same experiment and were processed in parallel.
- The localization of BCAS2 in the primordial, primary, secondary, and antral follicles. The lower panels show magnified views of the indicated regions (boxes above). Scale bars: 50 μm.
- Venn diagrams showing the overlap of abnormal AS genes and events in Hnrnpm and Bcas2 cKO oocytes. The overlaps of 98 common genes and 57 identical splicing events were both highly significant by hypergeometric test (p = 2e-27 and p = 4e-27, respectively).
- Sashimi plots showing the splicing patterns of Cenpf, Prc1, and Hdlbp in control, Hnrnpm cKO, and Bcas2 cKO oocytes.
- PCR validation of splicing events in (g–i). Schematic showing the alternatively spliced exons.
- Bar plots display the percent spliced- in (PSI) values, calculated as PSI = splice_in/(splice_in + splice_out). Two-sided Student's t-tests. Data are presented as mean ± SEM. n = 3. For Bcas2 ctrl and cKO group, p = 0.003473 (Cenpf), 0.000519 (Prc1), 0.007735 (Hdlbp). For Hnrnpm ctrl and cKO group, p = 0.009008 (Cenpf), 0.012079 (Prc1), 0.001602 (Hdlbp).
- RIP-PCR and qPCR analyses of hnRNPM and BCAS2-interacted transcripts in P21 ovaries. RNA immunoprecipitation followed using qPCR (RIP-qPCR) was performed to detect Cenpf, Prc1, and Hdlbp transcripts co-precipitated with anti-hnRNPM antibodies using anti-IgG as a negative control in P21 mouse ovaries. Two-sided Student's t-tests. Data are presented as mean ± SEM. n = 5.
- Western blotting confirmed the high knockdown efficiency of hnRNPM in HEK293T cells.
- RIP-PCR and qPCR analyses revealed BCAS2-RNA interactions in control and HNRNPM knockdown 293T cells. The association of CENPF, PRC1, and HDLBP transcripts with BCAS2 was examined using RIP-PCR (o) and qPCR (o). Two-sided Student's t-tests. Data are presented as mean ± SEM. n = 5. p = 0.000037 (PRC1). p < 0.000001 (CENPF, HDLBP).
2. Core Product Advantages of Absin abs580304
① Ultra-High Specificity for Precise Target Signal Capture
Abs580304 undergoes rigorous antigen screening and affinity purification processes, enabling specific recognition of target molecules while effectively avoiding cross-reactivity with homologous proteins. Even in complex clinical samples (such as tissue extracts or biological fluid specimens) or mixed cellular systems, it precisely captures target signals, preventing false-positive interference and ensuring accuracy in molecular expression detection.
② High Sensitivity for Low-Abundance Signal Detection
Utilizing advanced labeling technology and manufacturing processes, abs580304 features low detection limits and efficiently detects low-abundance target molecules. Whether in early-stage disease samples, limited cellular specimens, or scenarios involving subtle post-intervention expression changes, it accurately quantifies signal differences, providing a sensitive detection tool for mechanistic research.
③ Excellent Stability with High Batch-to-Batch Consistency
Strict adherence to ISO quality control standards, with comprehensive aseptic management from raw material procurement through production and packaging, ensures highly uniform activity, concentration, and specificity across batches. Multi-batch experiments and long-term tracking detection scenarios in research are supported with reproducible data, significantly reducing experimental errors caused by reagent variability.
④ Broad Compatibility Across Multiple Experimental Systems
Abs580304 flexibly adapts to various experimental techniques including Western blotting (WB), immunofluorescence (IF), immunohistochemistry (IHC), and flow cytometry, without requiring adjustment of core experimental conditions. It satisfies multi-dimensional validation needs in research, stably performing across molecular-level quantification, cellular-level localization, and tissue-level distribution observation, thereby enhancing experimental efficiency.
⑤ Ready-to-Use Formulation for Streamlined Experimental Workflow
Featuring a convenient ready-to-use formulation, abs580304 requires no complex preliminary dilution or activation steps—simply open and proceed with experiments. This design saves preparation time and reduces errors from improper handling, particularly suitable for high-throughput sample detection and multi-omics integrated analysis scenarios.
Core Contribution:Through these robust advantages, abs580304 successfully assisted the research team in precisely acquiring critical data on molecular expression levels, cellular localization, and signaling pathway activity, providing direct experimental evidence for core mechanistic validation and conclusion derivation, becoming the "core driving force" for the smooth advancement of this research.
IV. Brand Empowerment: Absin—Advancing Science Together, Building Innovation Pathways
Absin remains committed to addressing core needs in life science research and clinical translation, dedicated to providing high-quality, cost-effective experimental reagents and solutions. As a flagship product, abs580304 has been widely applied across multiple disciplines including tumor biology, clinical medicine, immunology, and neuroscience, contributing to thousands of published research articles and earning high recognition from researchers worldwide.
Moving forward, Absin will continue to deepen reagent research and development innovation, continuously optimizing product performance and expanding product portfolios. With more comprehensive product systems and more professional technical support, we empower researchers globally to explore the mysteries of life and overcome clinical challenges.
Contact Absin
Absin provides antibodies, proteins, ELISA kits, cell culture, detection kits, and other research reagents. If you have any product needs, please contact us.
| Absin Bioscience Inc. worldwide@absin.cn |
 Follow us on Facebook: Absin Bio Follow us on Facebook: Absin Bio |
