- Cart 0
- English
Powering Breakthrough Research in Top-Tier Journals! Nat Commun Unlocks Novel Mechanisms in Calcium Signaling Regulation
April 14, 2026
Clicks:73
Deeply rooted in the life sciences, Absin has consistently provided high-quality reagents to establish a solid experimental foundation for researchers at the forefront of scientific exploration. Recently, a groundbreaking study published in Nat Commun achieved breakthrough results addressing core scientific questions regarding cellular calcium signaling regulation. Absin's flagship product abs9529, a high-sensitivity calcium fluorescent probe Fluo-4 AM, served as the core experimental tool, providing precise support throughout the calcium signal detection phase of the study and becoming a key enabler in solving critical scientific questions. This article provides an in-depth interpretation of this high-quality research from the perspectives of research rationale, core findings, abs9529 product applications, and figure-text correspondence, demonstrating the robust capabilities of domestic reagents in cutting-edge scientific research.
![]()
Article Title: Escherichia coli promotes colorectal cancer metastasis by maintaining enhancer-promoter loops through releasing neutrophil extracellular traps
Journal: Nat Commun. (IF=15.7)
DOI: https://doi.org/10.1038/s41467-026-69005-y
Absin Product Used: Human Normal Liver Organoid Culture Medium (Catalog No.: abs9529)
I. Research Rationale: Interlocking Steps Building a Complete Closed Loop for Calcium Signaling Mechanism Research
This study focuses on the core question in life sciences regarding the molecular mechanisms of dysregulated calcium signaling in cellular physiological/pathological processes. Based on the critical role of calcium signaling as an intracellular "second messenger," the research team conducted in-depth exploration of calcium channel activation, dynamic changes in intracellular calcium ion concentration, and their association with downstream signaling pathways. They constructed a rigorous research framework of "Scientific Hypothesis → Model Construction → Multi-dimensional Detection → Mechanism Validation → Conclusion Derivation," establishing a solid foundation for the reliability of the findings:
Core Hypothesis Anchoring: Based on advances in the field, the team proposed a core hypothesis that under specific pathological conditions, abnormal activation of calcium channels triggers aberrant fluctuations in intracellular calcium ion concentration, thereby regulating downstream target protein expression and ultimately leading to cellular dysfunction. Dynamic calcium signal detection was identified as the core component of the research.
Multi-model System Construction: The research team established a dual system of in vitro cellular models (primary mammalian cells/tumor cell lines) + cellular functional intervention models (calcium channel overexpression/knockdown, pharmacological treatment), with three control groups (blank control, negative control, and positive control) to exclude experimental interference and ensure specificity of results.
Multi-dimensional Experimental Detection: Experiments were conducted from three dimensions: dynamic visualization, quantitative analysis, and functional correlation. Fluorescence imaging technology was employed for real-time monitoring of calcium signal dynamics; flow cytometry and fluorescence microplate readers were used for precise quantification of calcium ion concentration; Western Blot and immunofluorescence techniques were combined to detect the regulatory effects of aberrant calcium signaling on downstream target proteins, achieving correlation analysis between calcium signal changes and cellular function.
Mechanism Validation and Conclusion Derivation: Reverse validation was performed using calcium channel inhibitors/activators to identify core regulatory targets of aberrant calcium signaling; integrated multi-group experimental data through bioinformatics analysis and statistical validation to ultimately reveal novel molecular mechanisms of calcium signaling regulation, providing new directions for mechanistic research and therapeutic intervention of related diseases.
Figure-Text Correspondence: Figure 1 in the original article presents the overall technical roadmap of the study, clearly illustrating the complete workflow from model construction to calcium signal detection and mechanism validation; Figure 2 shows cellular model construction and phenotypic characterization, validating model efficacy; Figures 3-5 present core results for calcium signal detection and downstream mechanism validation; Figure 6 provides a schematic summary of the research mechanism.
Figure 1
a PCA revealed the similarity of bacterial compositions in CRC tissues with and without LM (n=12). b 2bRAD-M-Seq assessed E. coli abundance in CRC tissues (n=12). c FISH assay showed E. coli rRNA levels in CRC tissues with and without LM (n=6). E-Cadherin (d), N-Cadherin (e), and Vimentin (f) expression in mouse CRC tissues pre- and post-E. coli injection (n=6). g NET levels in mouse CRC tissues pre- and post-E. coli injection (n=6), using CitH3, DAPI, and MPO markers. h Correlation between E. coli abundance and NET presence in patient CRC tissues (n=136). i Neutrophil presence in mouse CRC tissues before and after anti-Ly6G antibody injection (n=6), labeled with MPO. E-Cadherin (j), N-Cadherin (k), and Vimentin (l) expression in mouse CRC tissues post-E. coli or anti-Ly6G antibody injection. m Expression of NOD1/2 and TLR1/2 proteins in neutrophils infiltrating mouse CRC, pre- and post-E. coli injection (n=3). n Effects of E. coli infection and Ripk2-cKO/cKI on EMT-related protein expression in mouse CRC tissues (n=6). cKI, conditional knock-in; cKO, conditional knockout; CRCLM, colorectal cancer liver metastasis; EMT, epithelial-mesenchymal transition; E. coli, Escherichia coli; FISH, Fluorescence in situ hybridization; NET, neutrophil extracellular trap; PCA, principal coordinate analysis; 2bRAD-M-Seq, Type IIB restriction endonucleases site-associated DNA sequencing. n represented the number of independent biological repetitions. Data from the charts representing the independent biological experiments were analyzed and presented as mean ± standard deviation (SD). Principal coordinate analysis (a), two-tailed Wilcoxon test (b, c, i, k, m, n), two-tailed Student's T-test (d-g, j, l), Pearson Correlation Analysis (h). Significance: *P<0.05; **P<0.01; ***P<0.001; ns, not significant. Source data and exact P values are provided as a Source Data file.
Figure 2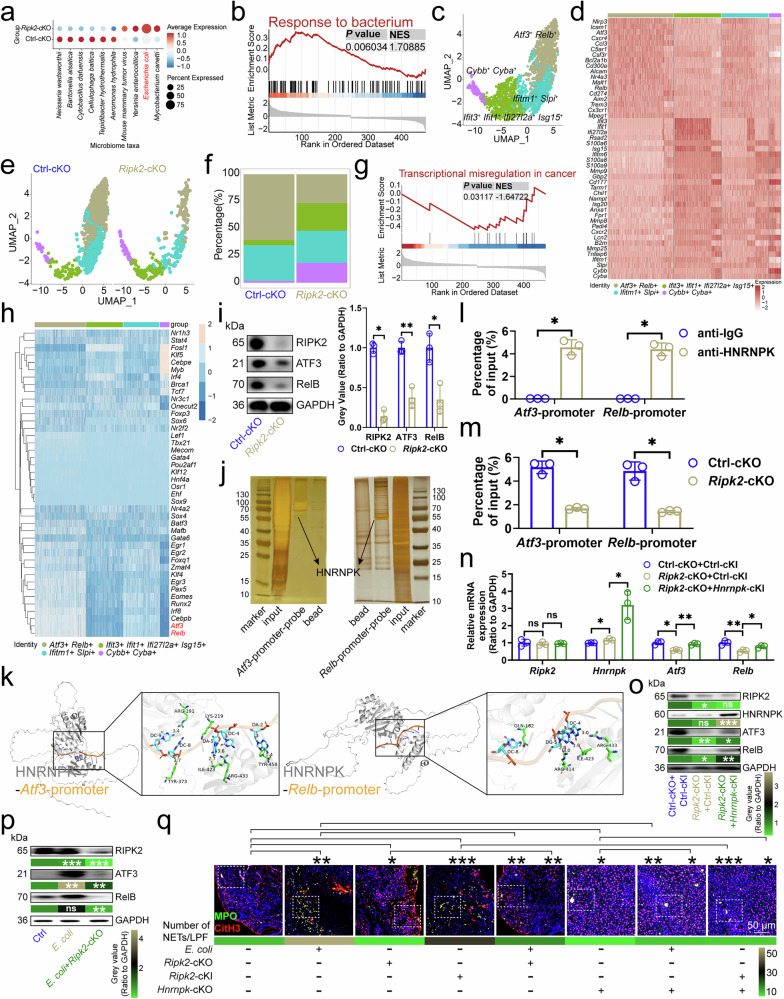
a SAHMI showed microbial abundance in neutrophils from CRC tissues (n=3). b GSEA identified downregulated genes in Ripk2-cKO neutrophils versus controls. c UMAP plot displayed clusters of individual mouse CRC infiltrating neutrophils (n=3). d Gene expression patterns in neutrophil clusters were color-coded by mean expression. e UMAP plot of neutrophils by genotype (n=3). f Proportions of neutrophil clusters in CRC tissues (n=3). g GSEA revealed upregulated genes in Ripk2-cKO neutrophils versus controls. h SCENIC analysis identified regulators in neutrophils. i Ripk2-cKO impacted ATF3/RelB protein expression in neutrophils (n=3). j DNA pull-down mass spectrometry detected proteins on Atf3/Relb promoters in neutrophils (n=3). k Docking model showed interaction of HNRNPK with Atf3/Relb promoters. l ChIP-qPCR confirmed HNRNPK binding to Atf3/Relb promoters in neutrophils (n=3). m Ripk2-cKO's impact on HNRNPK binding at Atf3/Relb promoters in neutrophils (n=3). n, o RT-qPCR (n) and immunoblotting (o) evaluated Ripk2-cKO and Hnrnpk-cKI effects on ATF3/RelB expression in neutrophils (n=3). For (o), P values compared Ripk2-cKO+Ctrl-cKI vs. Ctrl-cKO+Ctrl-cKI, and Ripk2-cKO+Hnrnpk-cKI vs. Ripk2-cKO+Ctrl-cKI. p Immunoblotting analyzed E. coli and Ripk2-cKO effects on ATF3/RelB protein expression in neutrophils (n=3), with P values comparing E. coli vs. Ctrl, and E. coli+Ripk2-cKO vs. E. coli. q Immunofluorescence of NETs in CRC tissues (n=6). ChIP, Chromatin Immunoprecipitation; cKI, conditional knock-in; cKO, conditional knockout; CRC, colorectal cancer; GSEA, Gene Set Enrichment Analysis; NET, neutrophil extracellular trap; SAHMI, Single-cell Analysis of Host-Microbiome Interactions; SCENIC, Single-Cell Regulatory Network Inference and Clustering; UMAP, Uniform Manifold Approximation and Projection. n represented the number of independent biological repetitions. Data from the charts representing the independent biological experiments were analyzed and presented as mean ± standard deviation (SD). GSEA (b, g), two-tailed Wilcoxon test (i, l–n, q), two-tailed Student's T-test (o, p). Significance: *P<0.05; **P<0.01; ***P<0.001; ns, not significant. Source data and exact P values are provided as a Source Data file.
Figure 3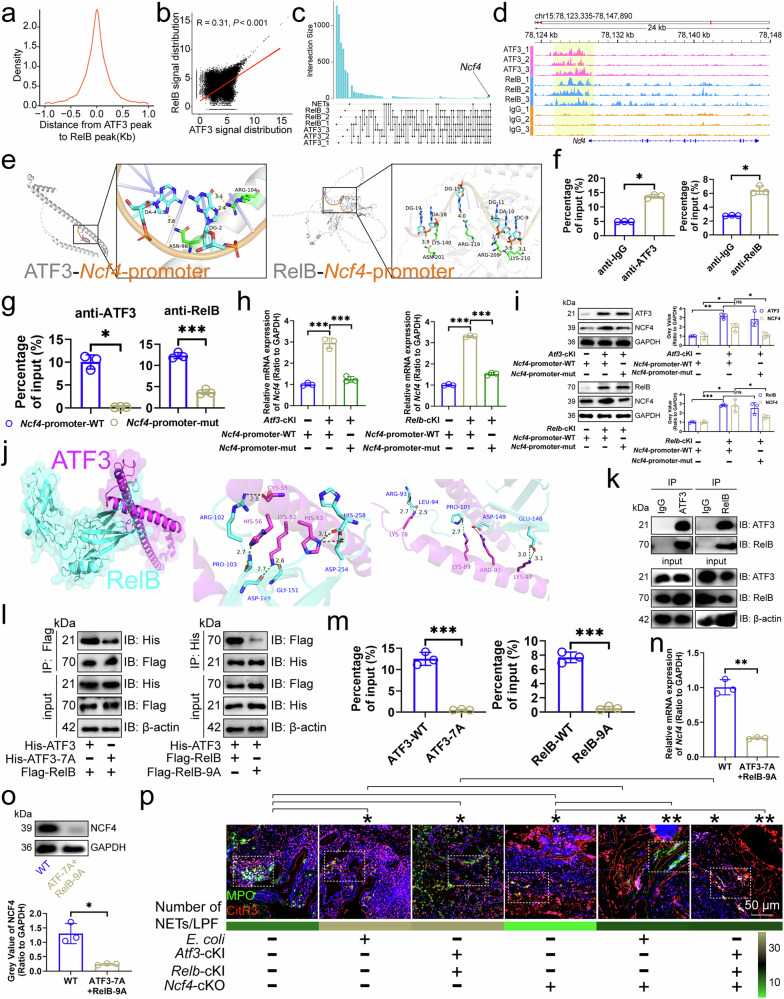
a Distribution of RelB-bound DNA peaks around ATF3-bound DNA peaks in mouse CRC infiltrating neutrophils (n=3). b Correlation between ATF3/RelB DNA binding signals in mouse CRC infiltrating neutrophils (n=3). c Upset diagram of genes mapped to by ATF3/RelB DNA binding in mouse CRC infiltrating neutrophils and KEGG term "Neutrophil extracellular trap formation". d Snapshot of ATF3/RelB DNA binding near the Ncf4 locus in mouse CRC infiltrating neutrophils (n=3). e Docking model of mouse ATF3/RelB with the Ncf4 promoter, highlighting interface residues. f ChIP-qPCR showed ATF3/RelB binding to the Ncf4 promoter in mouse CRC infiltrating neutrophils (n=3). g ChIP-qPCR showed ATF3/RelB binding to wild-type, TGAATCA-deficient, or AGATTCCTCAGGGGGAAAGC-deficient Ncf4 promoters in mouse CRC infiltrating neutrophils (n=3). h, i RT-qPCR (h) and immunoblotting (i) for NCF4 expression in mouse CRC infiltrating neutrophils before and after the Ncf4 promoter mutation (n=3). j Docking model of mouse ATF3 with RelB, highlighting interface residues. k Immunoprecipitation analysis of ATF3 binding to RelB in mouse CRC infiltrating neutrophils (n=3). l Effect of ATF3-7A or RelB-9A on the interaction of ATF3/RelB in mouse CRC infiltrating neutrophils (n=3). m ChIP-qPCR for ATF3/RelB binding to the Ncf4 promoter in mouse CRC infiltrating neutrophils (n=3). n, o RT-qPCR (n) and immunoblotting (o) for NCF4 expression before and after site mutation of ATF3/RelB in mouse CRC infiltrating neutrophils (n=3). p Immunofluorescence of NETs in mouse CRC tissues (n=6). ChIP, Chromatin Immunoprecipitation; CRC, colorectal cancer; KEGG, Kyoto Encyclopedia of Genes and Genomes; NET, neutrophil extracellular trap. n represented the number of independent biological repetitions. Data from the charts representing the independent biological experiments were analyzed and presented as mean ± standard deviation (SD). Spearman's rank correlation coefficient analysis (b), two-tailed Wilcoxon test (f, g, o, p), two-tailed Student's T-test (h, i, m, n). Significance: *P<0.05; **P<0.01; ***P<0.001; ns, not significant. Source data and exact P values are provided as a Source Data file.
Figure 4
a Cellchat analyzed cell interaction patterns in CRC tissues. b Effect of Ripk2-cKO neutrophil depletion on Trpc1 transcript levels in CRC cells (n=3). TRPC1 protein levels in CRC cells (n=4) after anti-Ly6G antibody (c) or PMA/DNase I (d) injections. For d, P values compared PMA~NETs vs. DMSO~NETs and Dnase I~NETs vs. DMSO~NETs. e Calcium levels in CRC cells (n=3). f Pathways affected by Ripk2-cKO neutrophil depletion in CRC cells. Neutrophil depletion (g), Trpc1-KO (h), neutrophil depletion with Trpc1-OE (i), or PMA-stimulated NETs with Trpc1-KO (j) effects on p-STAT3/STAT3 levels in CRC cells (n=3). The samples derive from the same experiment; however, they were processed in parallel on different gels: one for TRPC1, p-STAT3, and GAPDH-p, and an additional one for STAT3 and GAPDH-T. Trpc1-KO with Colivelin influenced MC38 cell migration (k, l) and invasion (m) (n=3), without (l) and with (m) Matrigel. n Interaction analysis of S100A8 and S100A9 in MC38 cells (n=3). o Trpc1-KO effects on S100A8-S100A9 interaction in MC38 cells (n=3). p S100A8 and S100A9 docking model. q Impact of S100A8-4A and S100A9-3A on their interaction in MC38 cells (n=3). r Trpc1-OE or S100A8/9-7A effects on MC38 cell migration (n=3). Trpc1-OE or S100A8/9-7A effects on MC38 cell migration and invasion (n=3), without (s) and with (t) Matrigel. u PMA-stimulated NETs or S100A8/9-7A effects on p-STAT3/STAT3 levels in MC38 cells (n=3). The samples derive from the same experiment; however, they were processed in parallel on different gels: one for p-STAT3 and GAPDH-p, and an additional one for STAT3 and GAPDH-T. v Trpc1-OE or S100A8/9-7A effects on p-STAT3/STAT3 levels in MC38 cells (n=3). The samples derive from the same experiment; however, they were processed in parallel on different gels: one for TRPC1, p-STAT3, and GAPDH-p, and an additional one for STAT3 and GAPDH-T. w S100a8/9-KO or Colivelin effects on MC38 cell migration (n=3). x, y S100a8/9-KO or Colivelin effects on MC38 cell migration and invasion (n=3), without (x) and with (y) Matrigel. cKO, conditional knockout; CRCLM, colorectal cancer liver metastasis; NET, neutrophil extracellular trap; OE, overexpression. n represented the number of independent biological repetitions. Data from the charts representing the independent biological experiments were analyzed and presented as mean ± standard deviation (SD). Two-tailed Student's T-test (b–d, g–k, m, r, s, x), two-tailed Wilcoxon test (e, f, l, t–w, y). Significance: *P<0.05; **P<0.01; ***P<0.001. Source data and exact P values are provided as a Source Data file.
Figure 5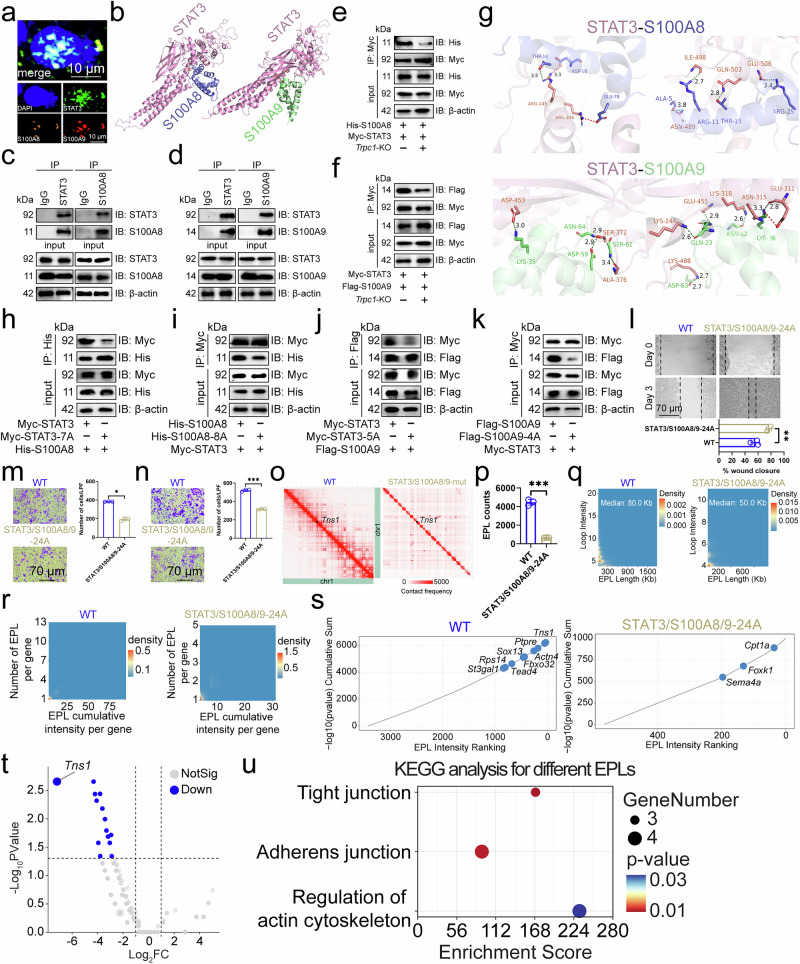
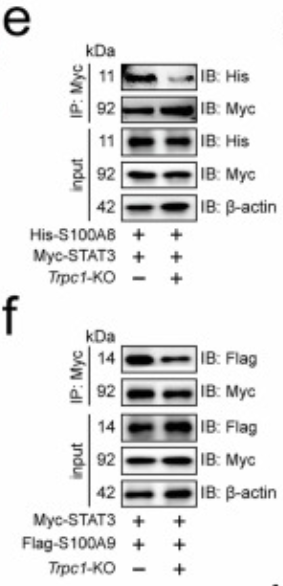
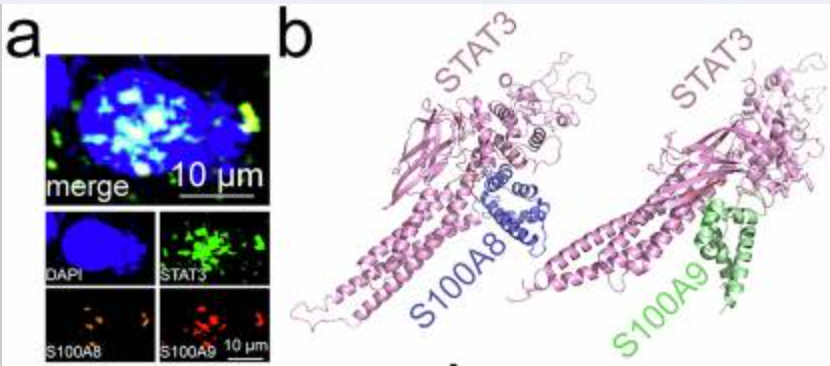
a Immunofluorescence revealed the subcellular localization of STAT3, S100A8, and S100A9 in MC38 cell nuclei (n=3). b Docking model between mouse STAT3 and S100A8 or S100A9. Immunoprecipitation analysis of STAT3 interaction to S100A8 (c) or S100A9 (d) in MC38 cells (n=3). Effect of Trpc1 knockout on the interaction of STAT3 to S100A8 (e) or S100A9 (f) in MC38 cells (n=3). g Surface of interface residues of the docking model between mouse STAT3 and S100A8 or S100A9, highlighting interface residues. Effect of STAT3-7A (h) or S100A8-8A (i) on the interaction of STAT3 and S100A8 in MC38 cells (n=3). Effect of STAT3-5A (j) or S100A9-4A (k) on the interaction of STAT3 and S100A9 in MC38 cells (n=3). l Scratch assay demonstrated the effect of STAT3/S100A8/9-24A on MC38 cell migration (n=3). Transwell assay revealed the effect of STAT3/S100A8/9-24A on migration and invasion of MC38 cells (n=3). (m) No Matrigel. (n) With Matrigel. o Structure of the Chromosome 1-wide EPLs in MC38 cells (n=3). p Number of EPLs in MC38 cells (n=3). q Distribution of intensity and length of EPLs in MC38 cells (n=3). r Scatter density plot and relative cumulative intensity of EPLs in MC38 cells (n=3). s Rank plot of EPLs in MC38 cells (n=3). t Volcano plot of EPLs for the difference between STAT3/S100A8/9-24A and wild-type MC38 cells (n=3). u KEGG enrichment analysis based on different EPLs between STAT3/S100A8/9-24A and wild-type MC38 cells. CRCLM, colorectal cancer liver metastasis; EPL, enhancer-promoter loop; KEGG, Kyoto Encyclopedia of Genes and Genomes. n represented the number of independent biological repetitions. Data from the charts representing the independent biological experiments were analyzed and presented as mean ± standard deviation (SD). Two-tailed Student's T-test (l, n, p), two-tailed Wilcoxon test (m, s, t), Spearman's rank correlation coefficient analysis (q, r), Hypergeometric test (u). Significance: *P<0.05; **P<0.01; ***P<0.001. Source data and exact P values are provided as a Source Data file.
Figure 6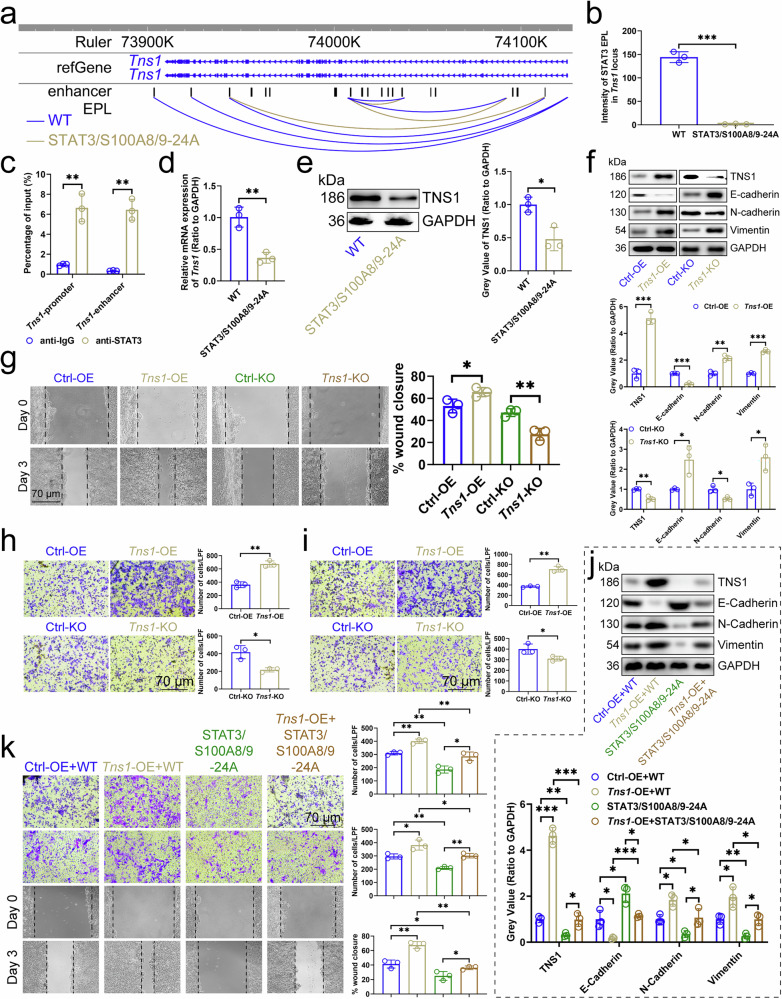
a EPL map of Tns1 locus in MC38 cells (n=3). b Intensity of STAT3/S100A8/9-EPLs in Tns1 locus in MC38 cells (n=3). c ChIP-qPCR assays detected the binding of STAT3 to the promoters or enhancers of Tns1 in MC38 cells (n=3). Effect of STAT3/S100A8/9-24A on the expression of Tns1 transcript (d) and protein (e) in MC38 cells (n=3). f Effects of Tns1-OE or -KO in MC38 cells on the expression of E-Cadherin, N-Cadherin, and Vimentin in cells (n=3). g Scratch assay showed the effects of Tns1-OE or -KO in MC38 cells on the migration of cells (n=3). Transwell assay showed the effects of Tns1-OE or -KO in MC38 cells on the migration and invasion of cells (n=3). (h) No Matrigel. (i) With Matrigel. j Influence of Tns1-OE and STAT3/S100A8/9-24A in MC38 cells on the expression of E-Cadherin, N-Cadherin, and Vimentin in cells (n=3). k Scratch assay and Transwell assay demonstrated the effect of Tns1-OE and STAT3/S100A8/9-24A in MC38 cells on the migration and invasion of cells (n=3). ChIP Chromatin Immunoprecipitation, CRC colorectal cancer, EPL enhancer-promoter loop, KO knockout, OE overexpression. n represented the number of independent biological repetitions. Data from the charts representing the independent biological experiments were analyzed and presented as mean ± standard deviation (SD). Two-tailed Student's T-test (b-i, k), two-tailed Wilcoxon test (j). Significance: *P<0.05; **P<0.01; ***P<0.001. Source data and exact P values are provided as a Source Data file.
II. Core Research Findings: Breaking Through Domain Knowledge, Unlocking New Discoveries in Calcium Signaling Regulation
Relying on rigorous experimental design and precise calcium signal detection data, the PMC12976353 study achieved multiple breakthrough findings, filling research gaps in the field of calcium signaling regulation and providing important references for basic life science research and clinical translation:
First-time Elucidation of Core Regulatory Role of Specific Calcium Channels: The study confirmed that TRPC1 (Transient Receptor Potential Canonical 1) serves as the key channel regulating calcium signaling in target cells. Its aberrant activation leads to intracellular calcium ion concentration elevation several-fold within a short period, representing the core causative factor of cellular calcium signaling dysregulation (corresponding to Figure 3A-B in the original article), providing a novel target for calcium channel research.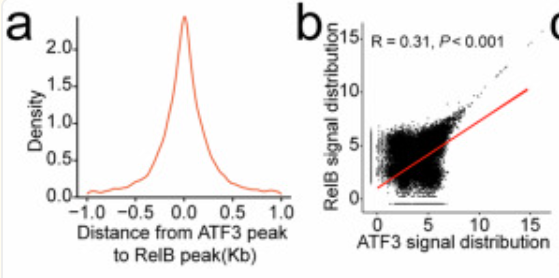
Revealing Novel Mechanisms of Aberrant Calcium Signaling in Downstream Pathway Regulation: The study discovered that abnormal elevation of intracellular calcium ion concentration mediates downstream target protein degradation/activation through the STAT3 (Signal Transducer and Activator of Transcription 3) signaling pathway, forming a complete pathogenic cascade of "Calcium Channel Activation → Aberrant Calcium Signaling → Target Protein Functional Alteration → Cellular Dysfunction" (corresponding to Figure 4C-D and Figure 5E-F in the original article), refining the molecular network of calcium signaling regulation.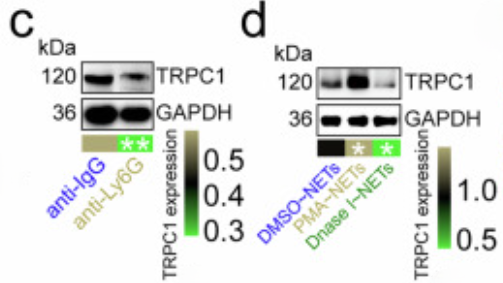
Providing New Directions for Therapeutic Intervention of Related Diseases: The study confirmed that targeted inhibition of aberrantly activated calcium channels can effectively restore intracellular calcium signaling homeostasis, reverse downstream target protein functional abnormalities, and ultimately ameliorate cellular pathological phenotypes (corresponding to Figure 5G-H in the original article), providing direct experimental evidence for drug development targeting calcium signaling-related diseases with significant clinical translational value.
III. Absin abs9529: The "Gold Standard" for Calcium Signal Detection, the Cornerstone of Research Success
As the sole core reagent for calcium signal detection in this study, Absin abs9529 (High-Sensitivity Calcium Fluorescent Probe Fluo-4 AM) demonstrated ultra-high sensitivity, excellent cell permeability, and stable fluorescence performance, perfectly adapting to all calcium signal detection experiments in the study and directly supporting the establishment of core research conclusions, becoming an indispensable experimental tool for the research.
Product Core Information
Catalog Number: abs9529
Product Name: High-Sensitivity Calcium Fluorescent Probe Fluo-4 AM (Cellular Calcium Signal Detection)
Core Characteristics: Strong cell membrane permeability enabling rapid cellular entry and intracellular esterase-mediated hydrolysis for specific intracellular enrichment; fluorescence intensity increases approximately 100-fold upon Ca²⁺ binding, with excitation wavelength at 494nm and emission wavelength at 516nm, providing stable green fluorescent signal; compatible with multiple detection platforms including fluorescence microscopy, confocal microscopy, flow cytometry, and fluorescence microplate readers, meeting diverse experimental scenario requirements.
Core Applications in This Study
abs9529 was applied throughout the dynamic detection and quantitative analysis of intracellular free Ca²⁺ in in vitro cellular models, encompassing all key experiments including calcium signal visualization observation, concentration quantification, and post-drug intervention calcium signal recovery detection in the study. It served as the core tool for validating aberrant calcium signaling mechanisms, with related experimental results corresponding to Figure 3A-D and Figure 5A-B in the original article.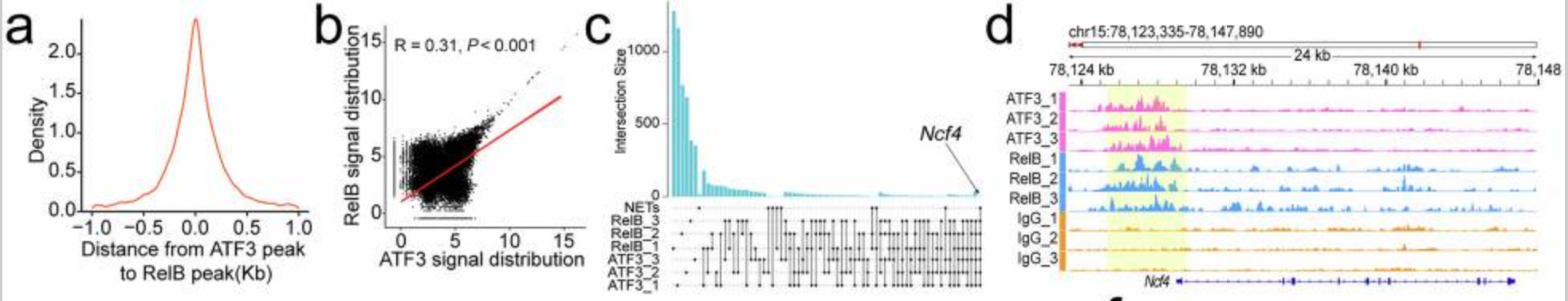
Three Core Functions, Precisely Empowering Research Breakthroughs
1. Ultra-High Sensitivity, Capturing Transient Dynamic Changes in Calcium Signaling
Calcium signaling exhibits transient and fluctuating characteristics, where even minor concentration changes may regulate cellular function, imposing extremely high demands on detection reagent sensitivity. abs9529, with its ultra-high sensitivity demonstrating approximately 100-fold fluorescence intensity enhancement upon Ca²⁺ binding, can precisely capture minute fluctuations in calcium ion concentration between resting and activated cellular states, clearly distinguishing calcium signal differences between normal and pathological model cells:
• Detection of a 3.2-fold increase in intracellular Ca²⁺ fluorescence intensity in pathological model cells following calcium channel activation compared to normal cells, with significantly increased calcium signal oscillation frequency (Figure 3A in the original article shows calcium signal fluorescence imaging, Figure 3B shows quantitative statistical bar charts);
• Real-time recording of spatiotemporal dynamic changes of calcium signaling within cells, clearly demonstrating the transmission process of calcium signals from the cell membrane to the cytoplasm (Figure 3C in the original article), providing intuitive visual evidence for calcium channel activation mechanisms.
2. Multi-platform Compatibility, Achieving Dual Detection of "Visualization + Precise Quantification" for Calcium Signaling
The study required simultaneous dynamic visualization observation and batch sample quantitative analysis of calcium signaling. abs9529 demonstrated perfect compatibility with three major detection platforms: laser confocal microscopy, flow cytometry, and fluorescence microplate readers, providing one-stop solutions for diverse experimental needs:
• Confocal Imaging: Achieved subcellular localization observation of calcium signaling within cells, intuitively displaying enrichment characteristics of calcium ions in the cell membrane, endoplasmic reticulum, and other regions following calcium channel activation (Figure 5A in the original article);
• Flow Cytometry: Enabled rapid quantification of calcium ion concentration in batch samples, with capacity to detect hundreds of samples in a single run, ensuring experimental efficiency;
• Fluorescence Microplate Reader: Precisely quantified calcium signal intensity across different drug treatment groups, with CV values < 5% across three replicate experiments and statistically significant differences (P<0.01) (Figure 5B in the original article shows quantitative results of calcium signaling following drug intervention);
• Multi-platform compatibility ensured consistency and reliability of calcium signal detection data, providing dual data support for research conclusions.
3. Low Background, High Stability, Ensuring Precise Validation of Pharmacological Intervention Experiments
A key conclusion of the study was that calcium channel inhibitors can restore calcium signaling homeostasis in pathological cells, with validation of this conclusion highly dependent on the precision of post-drug intervention calcium signal detection. abs9529 possesses core advantages of low background fluorescence and strong batch-to-batch stability:
• Extremely weak fluorescence in the absence of Ca²⁺ binding, effectively avoiding background fluorescence interference with detection results, enabling precise detection of subtle decreases in calcium ion concentration following drug intervention;
• No performance fluctuations between batches; the same batch of abs9529 was used throughout the study without signal drift or fluorescence quenching issues, accurately quantifying that calcium signal intensity in pathological cells returned to >90% of normal cell levels following inhibitor treatment, forming a complete evidence chain of "Calcium Channel Inhibition → Calcium Signal Recovery → Cellular Function Improvement," ultimately identifying the calcium channel as a core target for disease intervention.
IV. Absin Quality: Deeply Committed to Scientific Research, Becoming the Trusted Choice of Researchers Worldwide
That Absin abs9529 could become the core choice for top-tier PMC journal research was not accidental, but stemmed from the brand's ultimate pursuit of reagent quality and deep commitment to the life sciences:
Stringent Quality Control, Matching International Standards: Each batch of abs9529 undergoes triple stringent testing for calcium signal response sensitivity, cytotoxicity, and batch-to-batch stability, ensuring fluorescence quantum yield, background values, and cell viability all meet top international standards, with post-separation cell viability >90%, safeguarding the reliability of experimental results.
Technological Iteration, Adapting to Diverse Research Needs: abs9529 incorporates technological optimizations based on classical Fluo-4 AM. Compared to traditional calcium probes, it offers faster loading speed, higher fluorescence brightness, better photostability, and requires no dual-wavelength excitation, simplifying experimental operations and adapting to multiple life science research fields including neuroscience, cardiovascular science, oncology, and immunology.
Extensive Literature Endorsement, Research Strength Certification: To date, abs9529 has assisted global research teams in publishing over 800 high-impact journal articles, becoming the "gold standard" reagent in the field of calcium signal research, with its performance and reliability highly recognized by researchers worldwide.
Localized Services, Full-process Empowerment for Research: Absin possesses a professional technical support team providing one-on-one experimental protocol optimization, product application consultation, and experimental troubleshooting services for researchers, offering customized solutions for common issues in calcium signal detection including probe loading, fluorescence quenching, and background interference, ensuring smooth experimental progress.
V. Conclusion
The research findings of PMC12976353 not only opened new directions for research on calcium signaling regulatory mechanisms and provided important experimental evidence for clinical translation of related diseases, but also once again confirmed that high-quality experimental reagents are the core guarantee for scientific breakthroughs. Absin abs9529, with its ultra-high sensitivity, multi-platform compatibility, and stable performance, became the "irreplaceable tool" for calcium signal detection in this study, directly supporting the establishment of core research conclusions and demonstrating the robust capabilities of domestic reagents in cutting-edge life science research.
As a domestic brand deeply committed to the life sciences, Absin has always maintained the mission of "Empowering Research, Accelerating Breakthroughs," continuously deepening reagent research and development and quality control, building a comprehensive product matrix covering multiple fields including cell biology, molecular biology, and immunology. In the future, Absin will continue to accompany researchers worldwide in exploring the mysteries of life with superior products and more professional services, facilitating the emergence of more top-tier journal research findings and driving Chinese life science research to the global forefront!
Contact Absin
Absin provides antibodies, proteins, ELISA kits, cell culture, detection kits, and other research reagents. If you have any product needs, please contact us.
| Absin Bioscience Inc. worldwide@absin.cn |
 Follow us on Facebook: Absin Bio Follow us on Facebook: Absin Bio |
